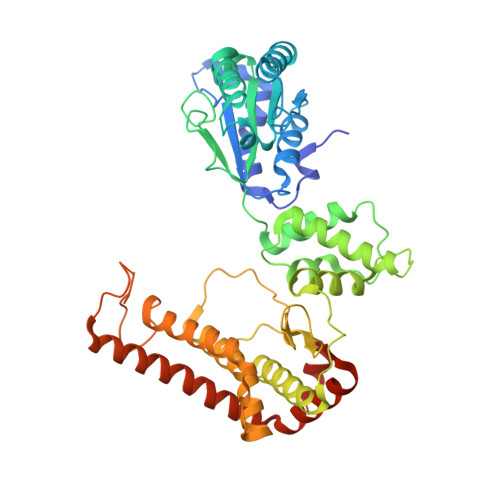
The crystal structure of apo-FtsH reveals domain movements necessary for substrate unfolding and translocation
Bieniossek, C., Niederhauser, B., Baumann, U.M.(2009) Proc Natl Acad Sci U S A 106: 21579-21584
- PubMed: 19955424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0910708106
- Primary Citation Related Structures:
3KDS - PubMed Abstract:
The hexameric membrane-spanning ATP-dependent metalloprotease FtsH is universally conserved in eubacteria, mitochondria, and chloroplasts, where it fulfills key functions in quality control and signaling. As a member of the self-compartmentalizing ATPases associated with various cellular activities (AAA+ proteases), FtsH converts the chemical energy stored in ATP via conformational rearrangements into a mechanical force that is used for substrate unfolding and translocation into the proteolytic chamber. The crystal structure of the ADP state of Thermotoga maritima FtsH showed a hexameric assembly consisting of a 6-fold symmetric protease disk and a 2-fold symmetric AAA ring. The 2.6 A resolution structure of the cytosolic region of apo-FtsH presented here reveals a new arrangement where the ATPase ring shows perfect 6-fold symmetry with the crucial pore residues lining an open circular entrance. Triggered by this conformational change, a substrate-binding edge beta strand appears within the proteolytic domain. Comparison of the apo- and ADP-bound structure visualizes an inward movement of the aromatic pore residues and generates a model of substrate translocation by AAA+ proteases. Furthermore, we demonstrate that mutation of a conserved glycine in the linker region inactivates FtsH.
- Department of Chemistry and Biochemistry, University of Bern, Freiestrasse 3, CH-3012 Bern, Switzerland.
Organizational Affiliation: