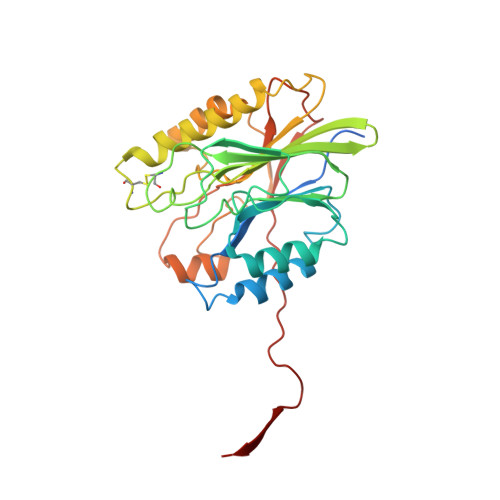
Structure of a mutant beta toxin from Staphylococcus aureus reveals domain swapping and conformational flexibility
Kruse, A.C., Huseby, M.J., Shi, K., Digre, J., Ohlendorf, D.H., Earhart, C.A.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 438-441
- PubMed: 21505235 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111005239
- Primary Citation Related Structures:
3K55 - PubMed Abstract:
The 3.35 Å resolution crystal structure of a mutant form of the staphylococcal sphingomyelinase β toxin in which a conserved hydrophobic β-hairpin has been deleted is reported. It is shown that this mutation induces domain swapping of a C-terminal β-strand, leading to the formation of dimers linked by a conformationally flexible hinge region. Eight dimers are seen in the asymmetric unit, exhibiting a broad spectrum of conformations trapped in place by intermolecular contacts within the crystal lattice. Furthermore, the 16 monomers within each asymmetric unit exhibit a remarkable heterogeneity in thermal factors, which can be accounted for by the varying degrees to which each monomer interacts with other molecules in the crystal. This structure provides a unique example of the challenges associated with crystallographic study of flexible proteins.
- Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, Minneapolis, MN 55455, USA.
Organizational Affiliation: