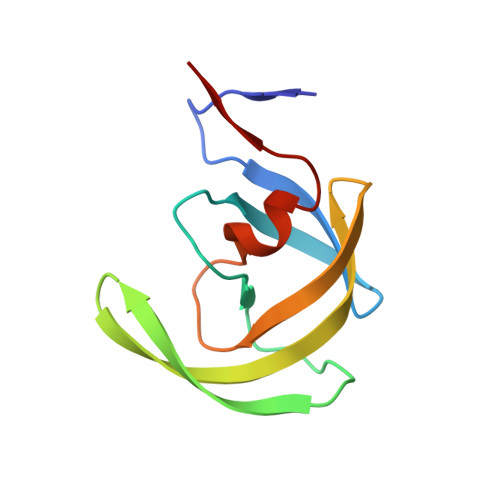
Carbamylation of N-terminal proline.
Olajuyigbe, F.M., Demitri, N., Ajele, J.O., Maurizio, E., Randaccio, L., Geremia, S.(2010) ACS Med Chem Lett 1: 254-257
- PubMed: 24900204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml100046d
- Primary Citation Related Structures:
3K4V - PubMed Abstract:
Protein carbamylation is of great concern both in vivo and in vitro. Here, we report the first structural characterization of a protein carbamylated at the N-terminal proline. The unexpected carbamylation of the α-amino group of the least reactive codified amino acid has been detected in high-resolution electron density maps of a new crystal form of the HIV-1 protease/saquinavir complex. The carbamyl group is found coplanar to the proline ring with a trans conformation. The reaction of N-terminal with cyanate ion derived from the chaotropic agent urea was confirmed by mass spectra analysis on protease single crystals. Implications of carbamylation process in vitro and in vivo are discussed.
- Department of Chemical Sciences ; Department of Biochemistry, Federal University of Technology, 340001 Akure, Nigeria.
Organizational Affiliation: