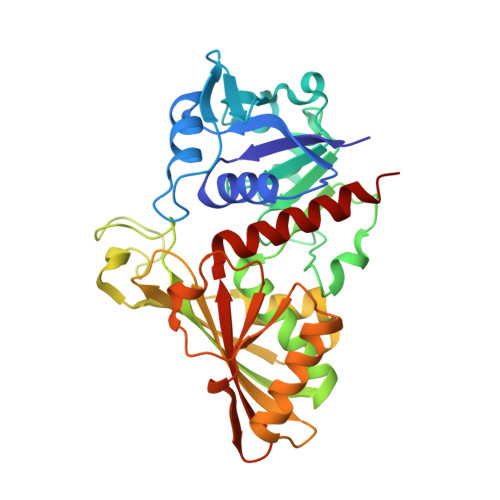
Structure of photosynthetic glyceraldehyde-3-phosphate dehydrogenase (isoform A4) from Arabidopsis thaliana in complex with NAD
Fermani, S., Sparla, F., Marri, L., Thumiger, A., Pupillo, P., Falini, G., Trost, P.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 621-626
- PubMed: 20516587 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309110013527
- Primary Citation Related Structures:
3K2B - PubMed Abstract:
The crystal structure of the A(4) isoform of photosynthetic glyceraldehyde-3-phosphate dehydrogenase (GAPDH) from Arabidopsis thaliana, expressed in recombinant form and complexed with NAD, is reported. The crystals, which were grown in 2.4 M ammonium sulfate and 0.1 M sodium citrate, belonged to space group I222. The asymmetric unit includes ten subunits, i.e. two independent tetramers plus a dimer that generates a third tetramer by a crystallographic symmetry operation. The crystal structure was solved by molecular replacement and refined to an R factor of 23.7% and an R(free) factor of 28.9% at 2.6 A resolution. In the final model, each subunit binds one NAD(+) molecule and two sulfates, which occupy the P(s) and the P(i) anion-binding sites. Detailed knowledge of this structure is instrumental for structural investigation of supramolecular complexes of A(4)-GAPDH, phosphoribulokinase and CP12, which are involved in the regulation of photosynthesis in the model plant A. thaliana.
- Department of Chemistry, University of Bologna, Via Selmi 2, 40126 Bologna, Italy.
Organizational Affiliation: