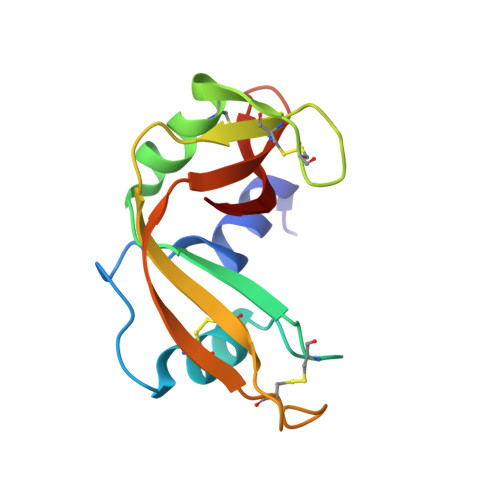
Structure of bovine pancreatic ribonuclease complexed with uridine 5'-monophosphate at 1.60 A resolution.
Larson, S.B., Day, J.S., Nguyen, C., Cudney, R., McPherson, A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 113-120
- PubMed: 20124705 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S174430910905194X
- Primary Citation Related Structures:
3JW1 - PubMed Abstract:
Bovine pancreatic ribonuclease A (RNase A) was crystallized from a mixture of small molecules containing basic fuchsin, tobramycin and uridine 5'-monophosphate (U5P). Solution of the crystal structure revealed that the enzyme was selectively bound to U5P, with the pyrimidine ring of U5P residing in the pyrimidine-binding site at Thr45. The structure was refined to an R factor of 0.197 and an R(free) of 0.253.
- Department of Molecular Biology and Biochemistry, The University of California, Irvine, CA 92697-3900, USA.
Organizational Affiliation: