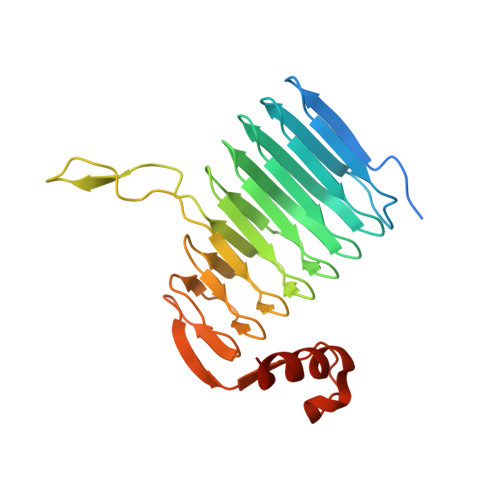
Crystal structure analysis of the polysialic acid specific O-acetyltransferase NeuO
Schulz, E.C., Bergfeld, A.K., Ficner, R., Muhlenhoff, M.(2011) PLoS One 6: e17403-e17403
- PubMed: 21390252 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0017403
- Primary Citation Related Structures:
3JQY - PubMed Abstract:
The major virulence factor of the neuroinvasive pathogen Escherichia coli K1 is the K1 capsule composed of α2,8-linked polysialic acid (polySia). K1 strains harboring the CUS-3 prophage modify their capsular polysaccharide by phase-variable O-acetylation, a step that is associated with increased virulence. Here we present the crystal structure of the prophage-encoded polysialate O-acetyltransferase NeuO. The homotrimeric enzyme belongs to the left-handed β-helix (LβH) family of acyltransferases and is characterized by an unusual funnel-shaped outline. Comparison with other members of the LβH family allowed the identification of active site residues and proposal of a catalytic mechanism and highlighted structural characteristics of polySia specific O-acetyltransferases. As a unique feature of NeuO, the enzymatic activity linearly increases with the length of the N-terminal poly-ψ-domain which is composed of a variable number of tandem copies of an RLKTQDS heptad. Since the poly-ψ-domain was not resolved in the crystal structure it is assumed to be unfolded in the apo-enzyme.
- Abteilung für Molekulare Strukturbiologie, Institut für Mikrobiologie und Genetik, Georg-August-Universität Göttingen, Göttingen, Germany.
Organizational Affiliation: