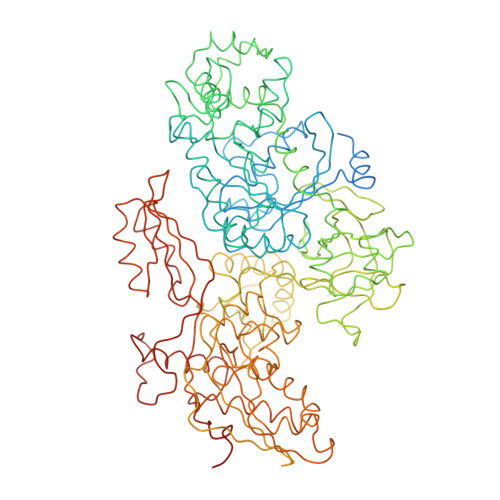
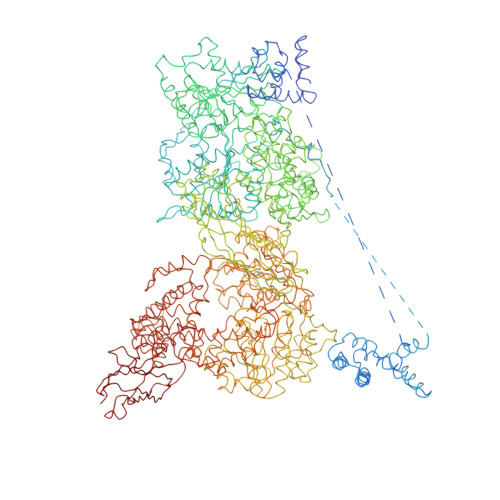
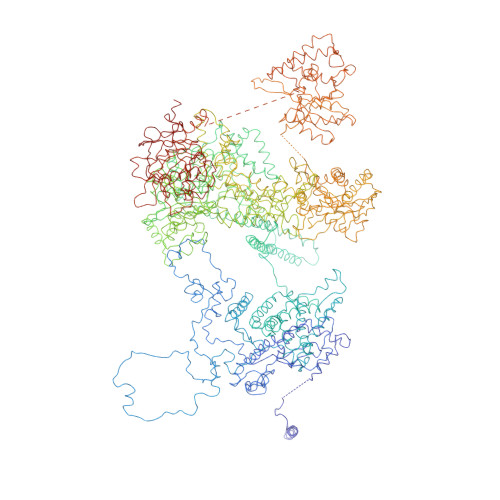
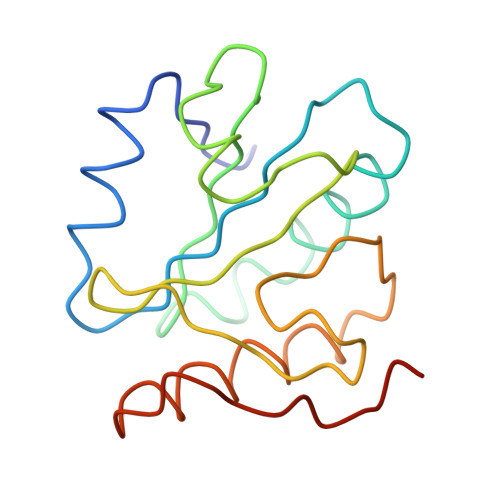
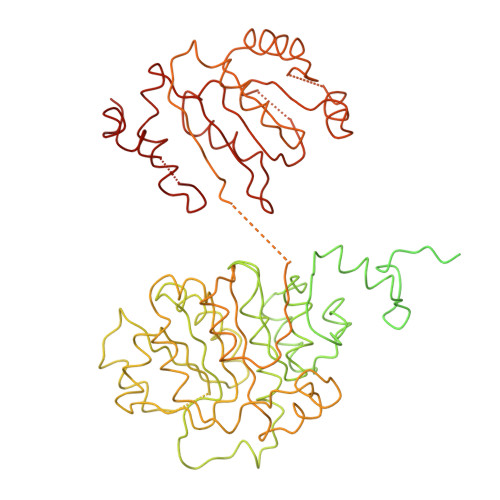
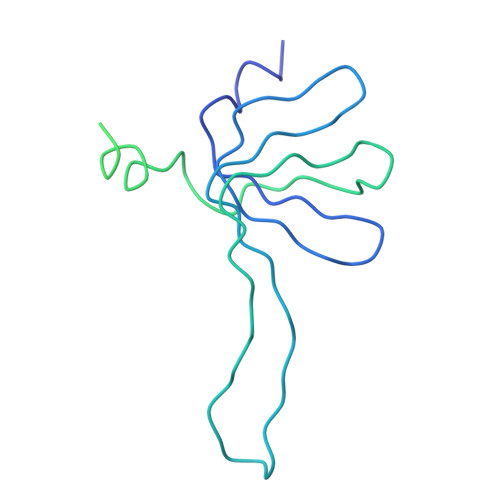
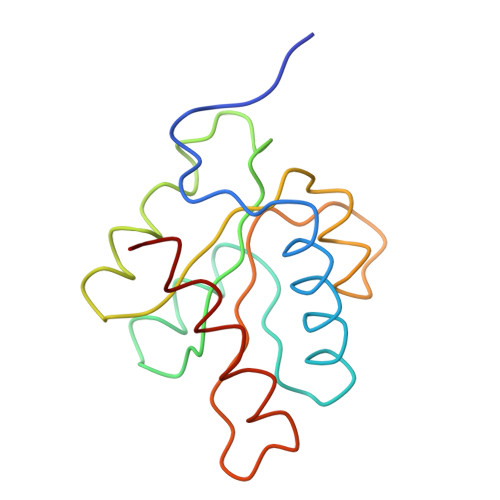
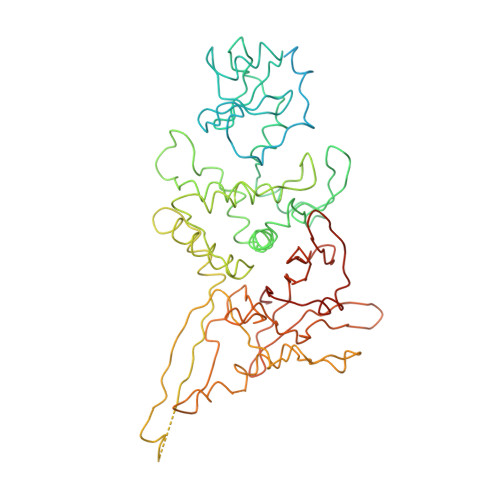
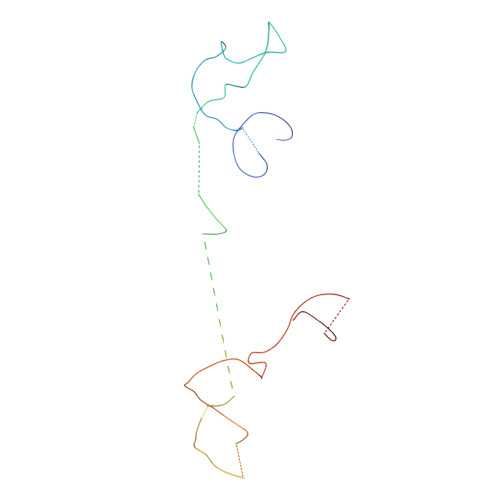
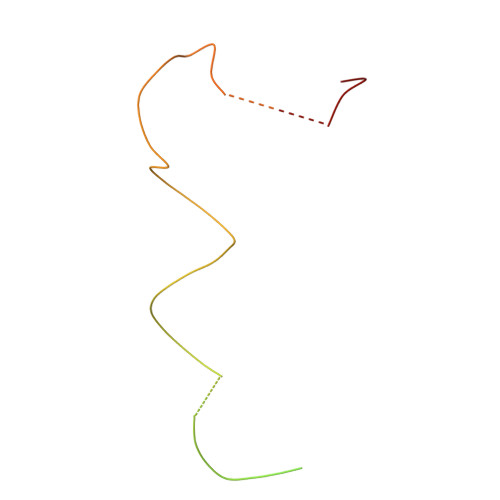
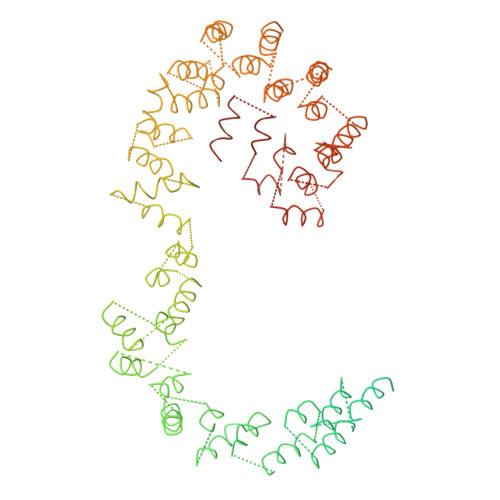Molecular architecture of the human U4/U6.U5 tri-snRNP.
Agafonov, D.E., Kastner, B., Dybkov, O., Hofele, R.V., Liu, W.T., Urlaub, H., Luhrmann, R., Stark, H.(2016) Science 351: 1416-1420
- PubMed: 26912367 Search on PubMed
- DOI: https://doi.org/10.1126/science.aad2085
- Primary Citation Related Structures:
3JCR - PubMed Abstract:
The U4/U6.U5 triple small nuclear ribonucleoprotein (tri-snRNP) is a major spliceosome building block. We obtained a three-dimensional structure of the 1.8-megadalton human tri-snRNP at a resolution of 7 angstroms using single-particle cryo-electron microscopy (cryo-EM). We fit all known high-resolution structures of tri-snRNP components into the EM density map and validated them by protein cross-linking. Our model reveals how the spatial organization of Brr2 RNA helicase prevents premature U4/U6 RNA unwinding in isolated human tri-snRNPs and how the ubiquitin C-terminal hydrolase-like protein Sad1 likely tethers the helicase Brr2 to its preactivation position. Comparison of our model with cryo-EM three-dimensional structures of the Saccharomyces cerevisiae tri-snRNP and Schizosaccharomyces pombe spliceosome indicates that Brr2 undergoes a marked conformational change during spliceosome activation, and that the scaffolding protein Prp8 is also rearranged to accommodate the spliceosome's catalytic RNA network.
- Department of Cellular Biochemistry, Max Planck Institute for Biophysical Chemistry, D-37077 Göttingen, Germany.
Organizational Affiliation: