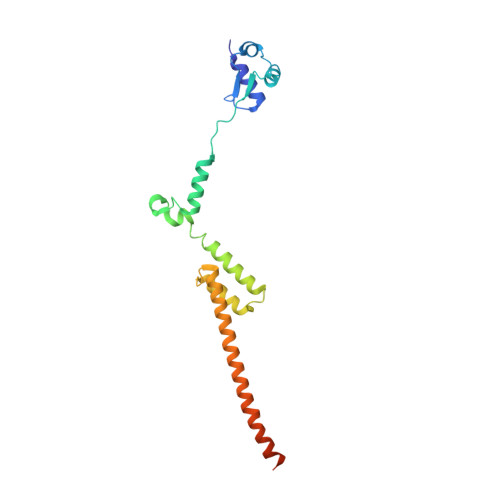
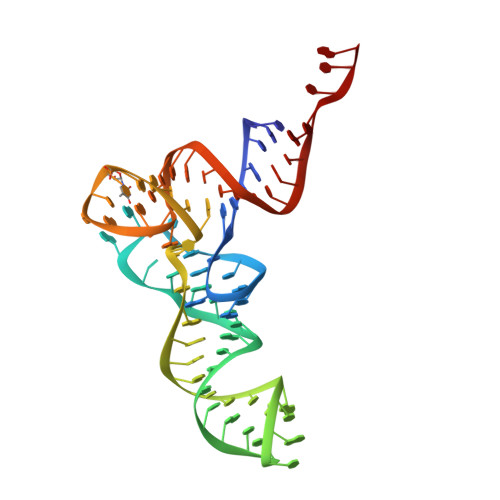
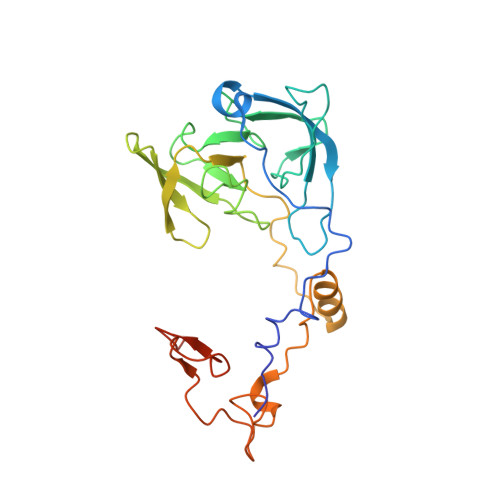
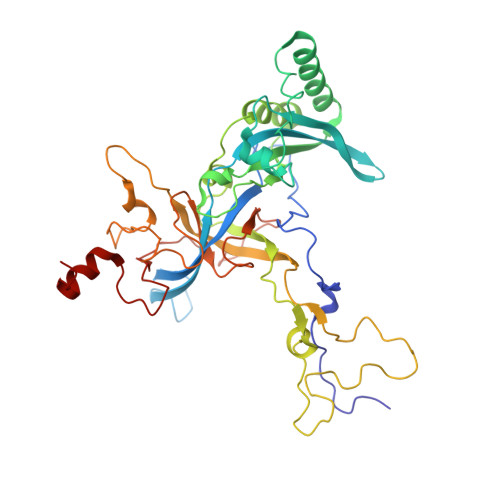
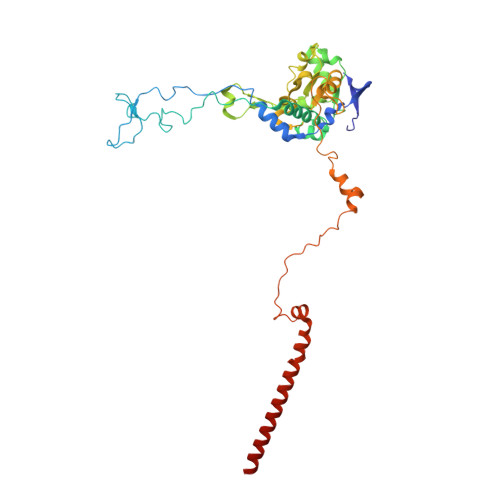
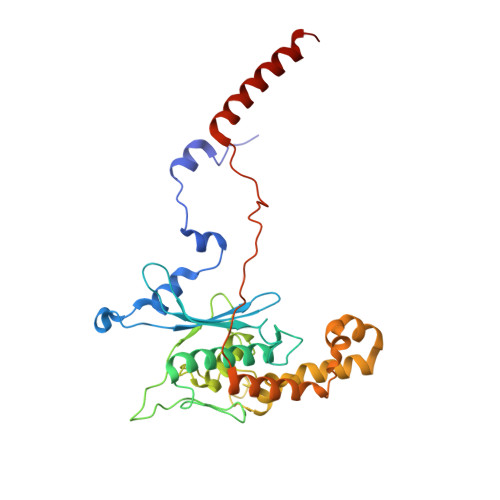
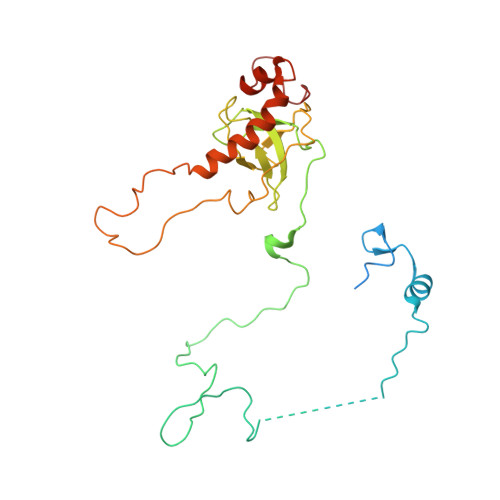
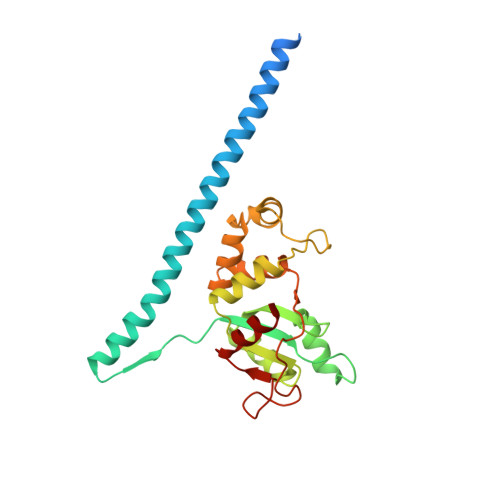
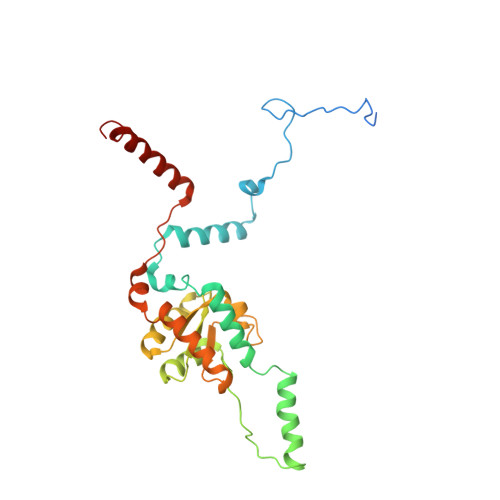
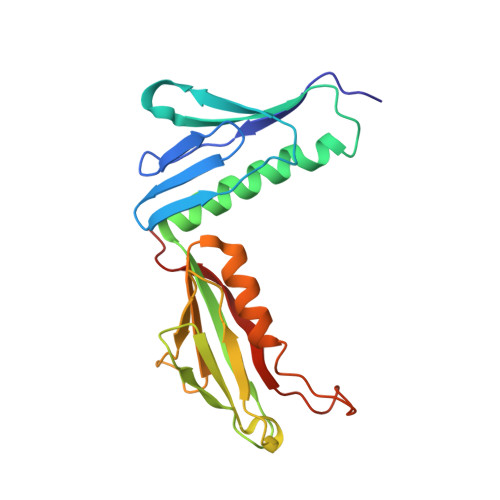
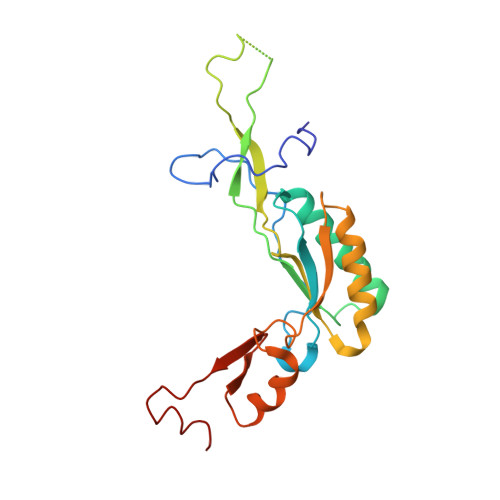
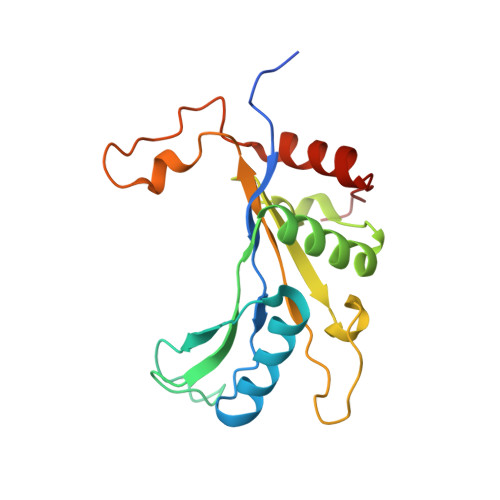
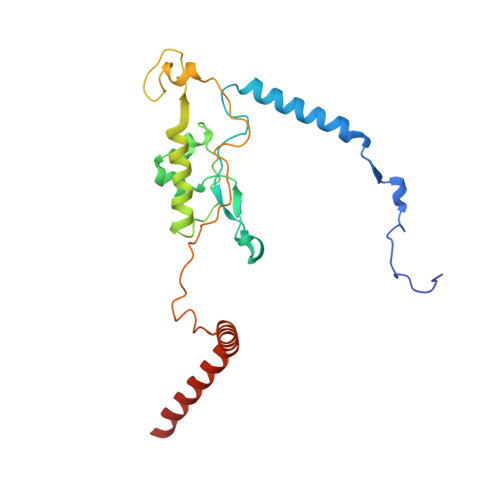
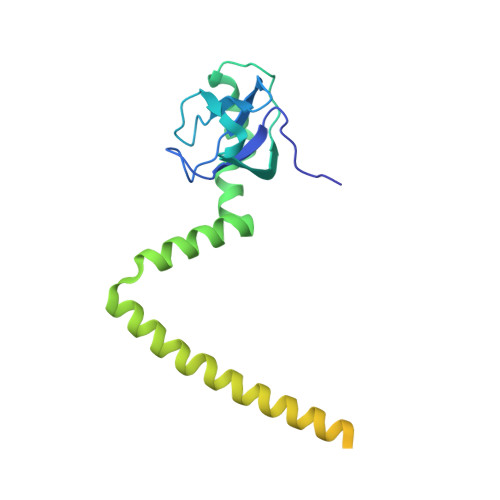
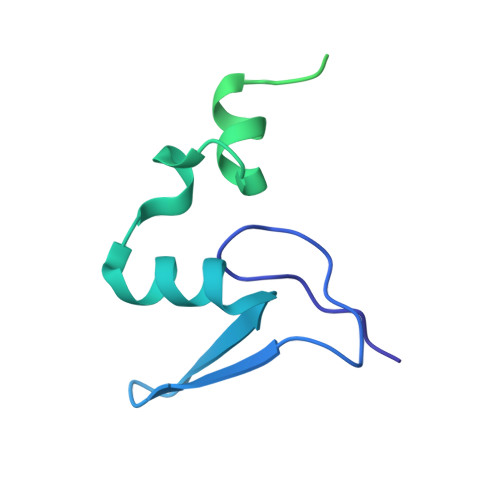
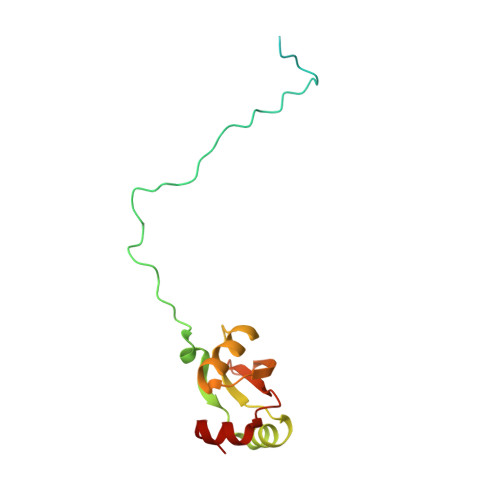
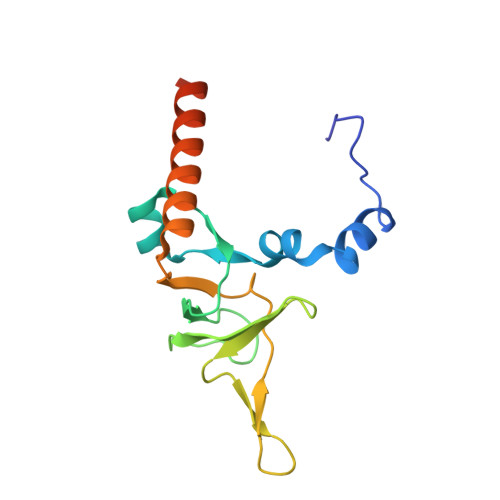
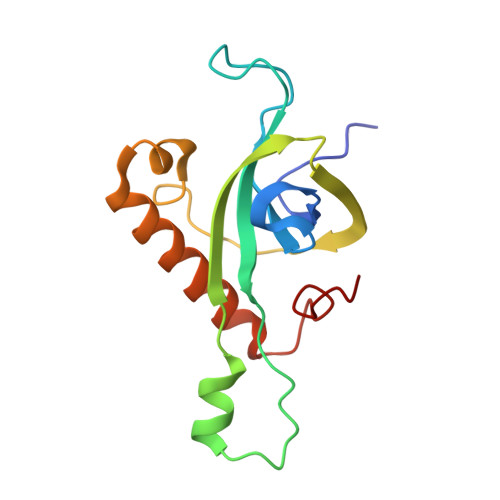
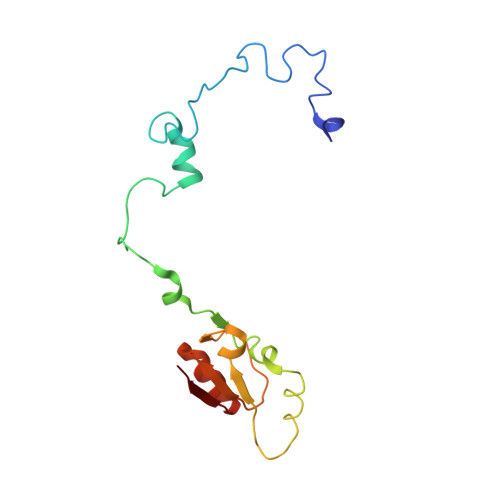
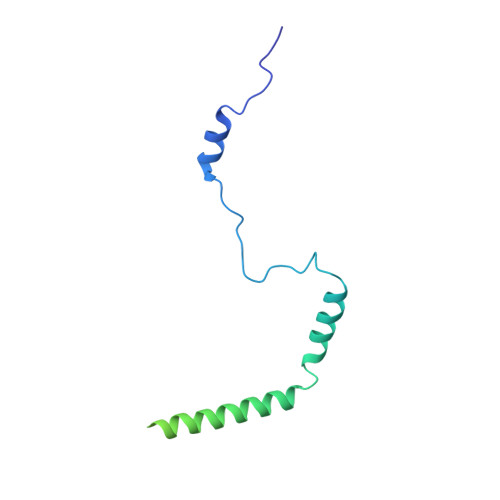
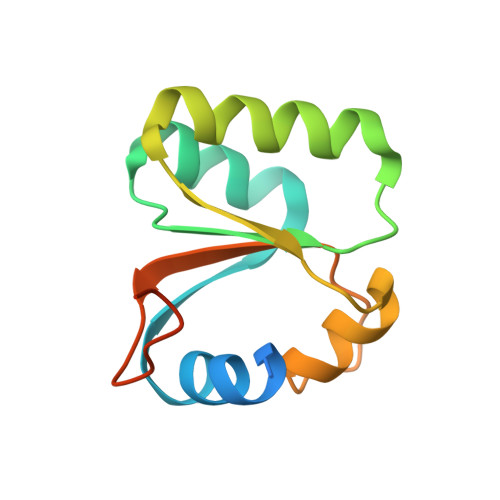
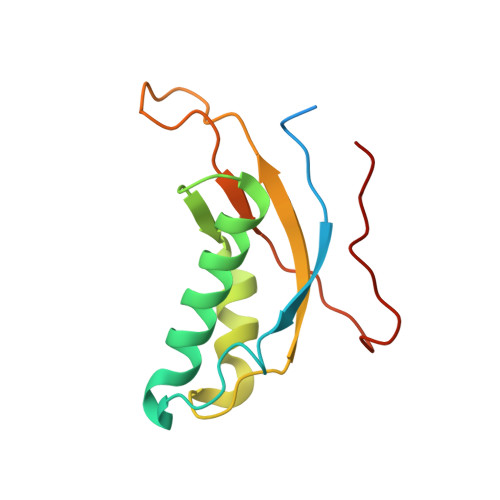
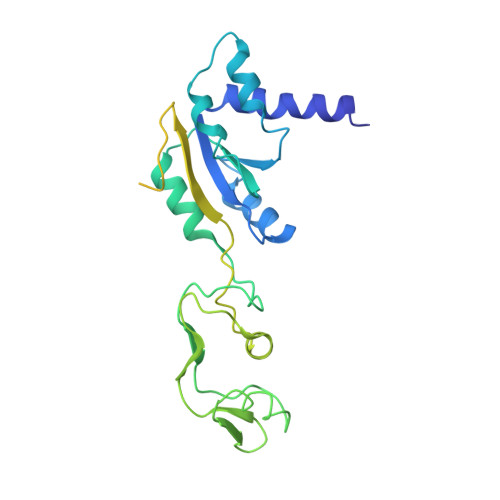
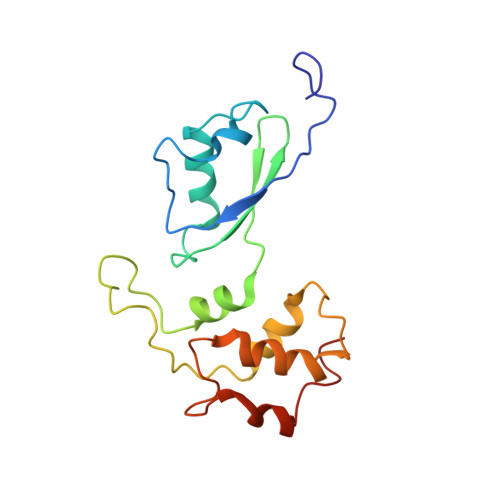
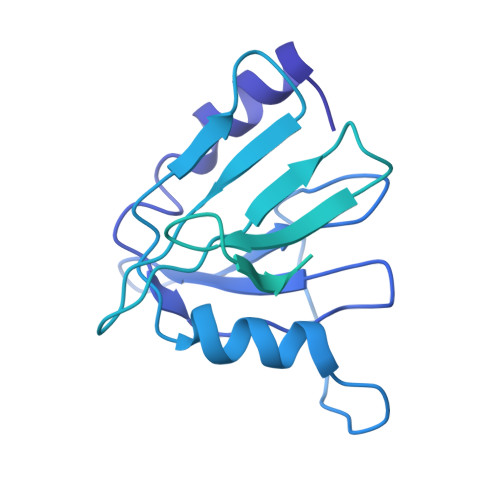
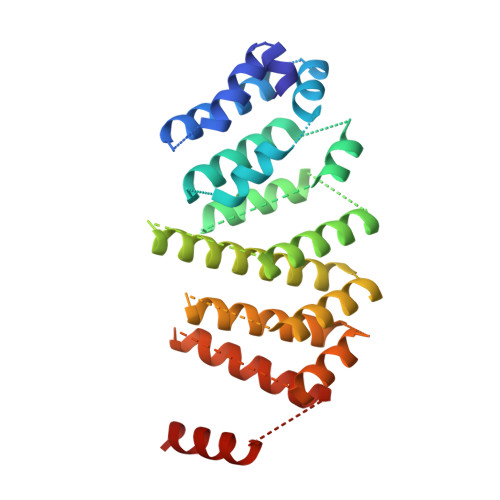
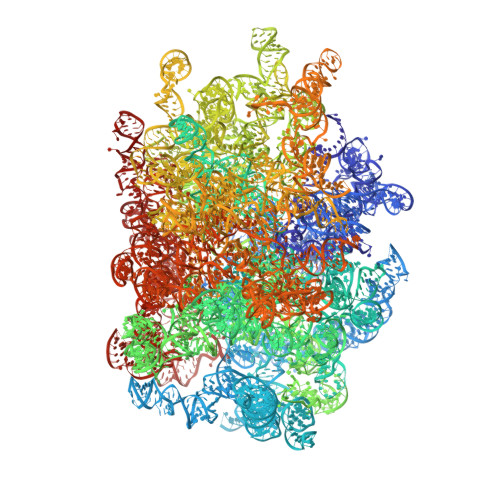
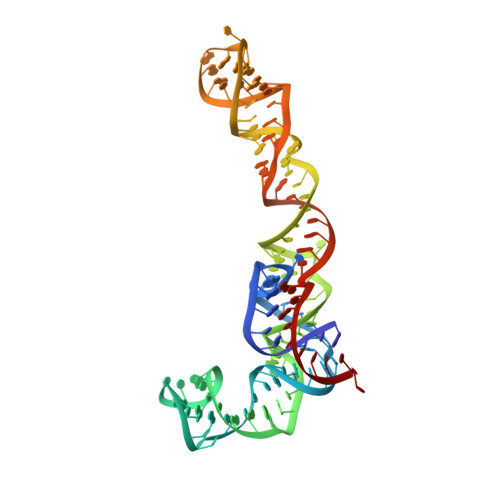
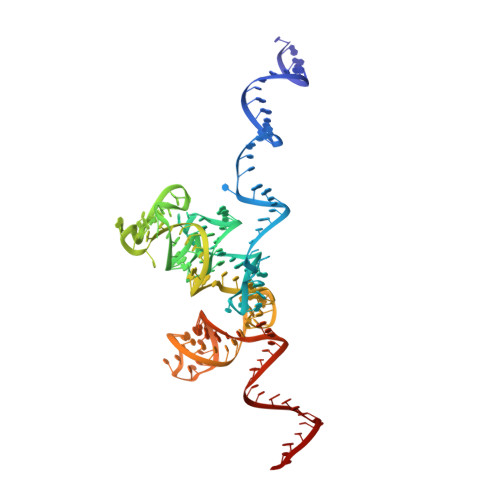
Structure and Assembly Pathway of the Ribosome Quality Control Complex.
Shao, S., Brown, A., Santhanam, B., Hegde, R.S.(2015) Mol Cell 57: 433-444
- PubMed: 25578875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2014.12.015
- Primary Citation Related Structures:
3J92 - PubMed Abstract:
During ribosome-associated quality control, stalled ribosomes are split into subunits and the 60S-housed nascent polypeptides are poly-ubiquitinated by Listerin. How this low-abundance ubiquitin ligase targets rare stall-generated 60S among numerous empty 60S is unknown. Here, we show that Listerin specificity for nascent chain-60S complexes depends on nuclear export mediator factor (NEMF). The 3.6 Å cryo-EM structure of a nascent chain-containing 60S-Listerin-NEMF complex revealed that NEMF makes multiple simultaneous contacts with 60S and peptidyl-tRNA to sense nascent chain occupancy. Structural and mutational analyses showed that ribosome-bound NEMF recruits and stabilizes Listerin's N-terminal domain, while Listerin's C-terminal RWD domain directly contacts the ribosome to position the adjacent ligase domain near the nascent polypeptide exit tunnel. Thus, highly specific nascent chain targeting by Listerin is imparted by the avidity gained from a multivalent network of context-specific individually weak interactions, highlighting a new principle of client recognition during protein quality control.
- MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge, CB2 0QH, UK.
Organizational Affiliation: