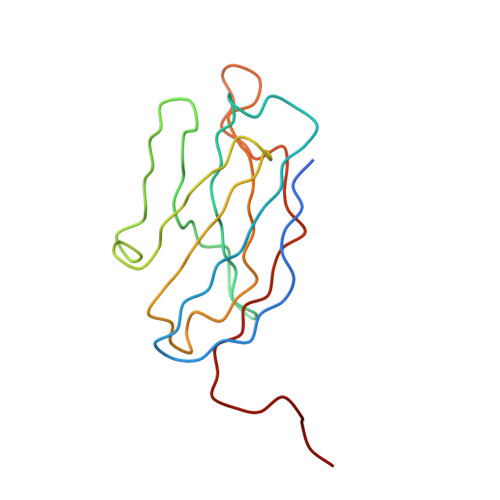
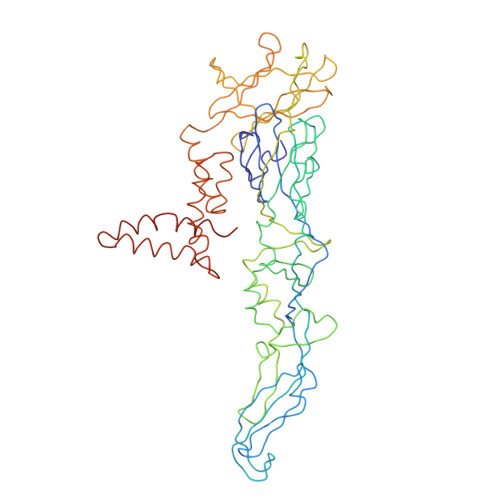
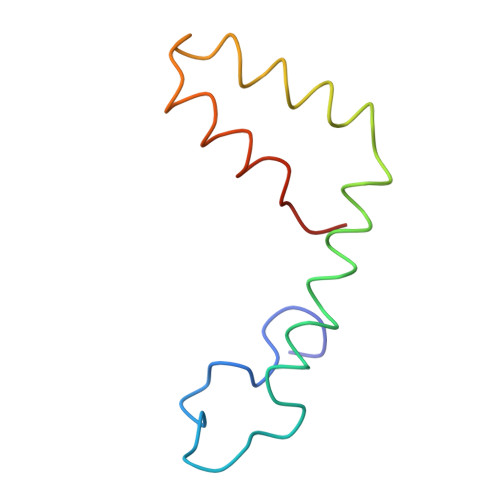
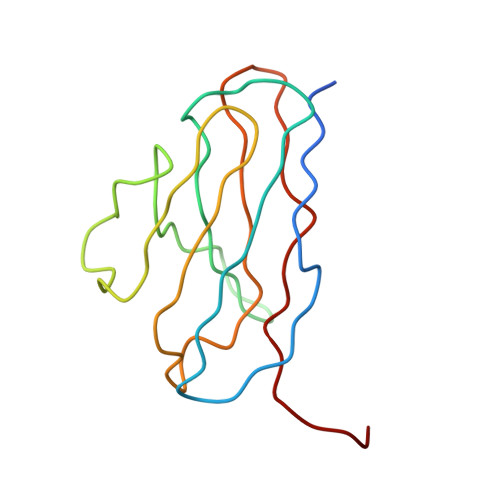
A highly potent human antibody neutralizes dengue virus serotype 3 by binding across three surface proteins.
Fibriansah, G., Tan, J.L., Smith, S.A., de Alwis, R., Ng, T.S., Kostyuchenko, V.A., Jadi, R.S., Kukkaro, P., de Silva, A.M., Crowe, J.E., Lok, S.M.(2015) Nat Commun 6: 6341-6341
- PubMed: 25698059 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms7341
- Primary Citation Related Structures:
3J6S, 3J6T, 3J6U - PubMed Abstract:
Dengue virus (DENV) infects ~400 million people annually. There is no licensed vaccine or therapeutic drug. Only a small fraction of the total DENV-specific antibodies in a naturally occurring dengue infection consists of highly neutralizing antibodies. Here we show that the DENV-specific human monoclonal antibody 5J7 is exceptionally potent, neutralizing 50% of virus at nanogram-range antibody concentration. The 9 Å resolution cryo-electron microscopy structure of the Fab 5J7-DENV complex shows that a single Fab molecule binds across three envelope proteins and engages three functionally important domains, each from a different envelope protein. These domains are critical for receptor binding and fusion to the endosomal membrane. The ability to bind to multiple domains allows the antibody to fully coat the virus surface with only 60 copies of Fab, that is, half the amount compared with other potent antibodies. Our study reveals a highly efficient and unusual mechanism of molecular recognition by an antibody.
- 1] Program in Emerging Infectious Diseases, Duke-NUS Graduate Medical School, 8 College Road, Singapore 169857, Singapore [2] Centre for BioImaging Sciences, National University of Singapore, 14 Science Drive 4, Singapore 117557, Singapore.
Organizational Affiliation: