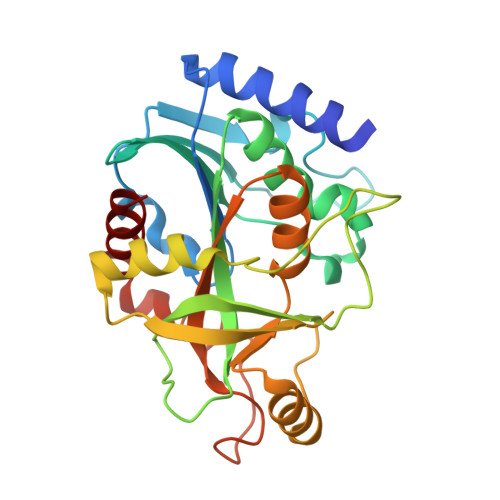
Crystal structure and molecular dynamics studies of purine nucleoside phosphorylase from Mycobacterium tuberculosis associated with acyclovir.
Caceres, R.A., Timmers, L.F., Ducati, R.G., da Silva, D.O., Basso, L.A., de Azevedo Jr., W.F., Santos, D.S.(2012) Biochimie 94: 155-165
- PubMed: 22033138 Search on PubMed
- DOI: https://doi.org/10.1016/j.biochi.2011.10.003
- Primary Citation Related Structures:
3IX2 - PubMed Abstract:
Consumption has been a scourge of mankind since ancient times. This illness has charged a high price to human lives. Many efforts have been made to defeat Mycobacterium tuberculosis (Mt). The M. tuberculosis purine nucleoside phosphorylase (MtPNP) is considered an interesting target to pursuit new potential inhibitors, inasmuch it belongs to the purine salvage pathway and its activity might be involved in the mycobacterial latency process. Here we present the MtPNP crystallographic structure associated with acyclovir and phosphate (MtPNP:ACY:PO(4)) at 2.10 Å resolution. Molecular dynamics simulations were carried out in order to dissect MtPNP:ACY:PO(4) structural features, and the influence of the ligand in the binding pocket stability. Our results revealed that the ligand leads to active site lost of stability, in agreement with experimental results, which demonstrate a considerable inhibitory activity against MtPNP (K(i) = 150 nM). Furthermore, we observed that some residues which are important in the proper ligand's anchor into the human homologous enzyme do not present the same importance to MtPNP. Therewithal, these findings contribute to the search of new specific inhibitors for MtPNP, since peculiarities between the mycobacterial and human enzyme binding sites have been identified, making a structural-based drug design feasible.
- Faculdade de Biociências, Laboratório de Bioquímica Estrutural, Pontifícia Universidade Católica do Rio Grande do Sul, Porto Alegre - RS, Brazil.
Organizational Affiliation: