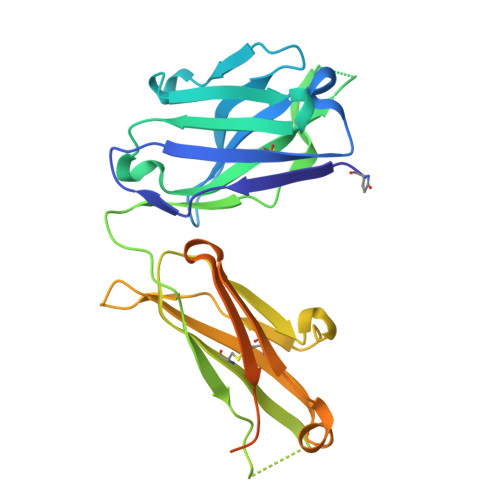
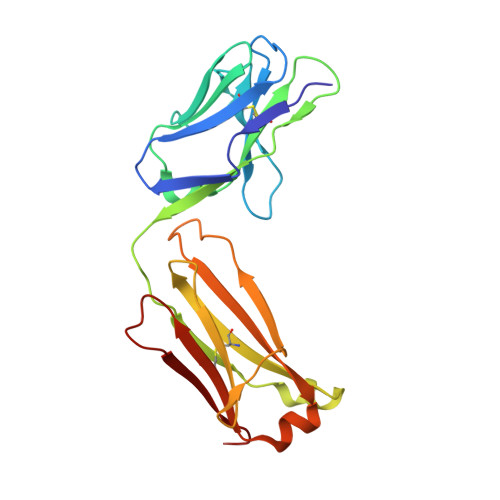
Crystal structure of an anti-ganglioside antibody, and modelling of the functional mimicry of its NeuGc-GM3 antigen by an anti-idiotypic antibody.
Talavera, A., Eriksson, A., Okvist, M., Lopez-Requena, A., Fernandez-Marrero, Y., Perez, R., Moreno, E., Krengel, U.(2009) Mol Immunol 46: 3466-3475
- PubMed: 19748674 Search on PubMed
- DOI: https://doi.org/10.1016/j.molimm.2009.07.032
- Primary Citation Related Structures:
3IU4 - PubMed Abstract:
N-Glycolylated (NeuGc) gangliosides are tumor-specific antigens and as such represent attractive targets for cancer immunotherapy. The chimeric antibody chP3 selectively recognizes a broad variety of NeuGc gangliosides, showing no cross-reactivity to the highly similar N-acetylated (NeuAc) gangliosides that are common cellular antigens in humans. Here, we report the crystal structure of the chP3 Fab and its computer-docking model with the trisaccharide NeuGcalpha3Galbeta4Glcbeta, which represents the carbohydrate moiety of the tumor-antigen NeuGc-GM3. The interaction involves only the heavy chain of the chP3 antibody. The modelled complex is consistent with all available experimental data and shows good surface complementarity. The negatively charged sialic acid residue NeuGc is buried in a pocket flanked by two arginine residues, VH Arg31 and VH Arg100A. We have further investigated the interaction of chP3 with its anti-idiotypic antibody, 1E10 (also known as Racotumomab), currently in clinical trials as a cancer vaccine. While many of the chP3 residues predicted to interact with the NeuGc ganglioside also feature prominently in the modelled complex of chP3 and 1E10, we do not observe structural mimicry. Rather, we suspect that the anti-idiotype 1E10 may serve as an imprint of the structural characteristics of the chP3 idiotype and, consequently, give rise to antibodies with P3-like properties upon immunization.
- Center of Molecular Immunology, Havana, Cuba.
Organizational Affiliation: