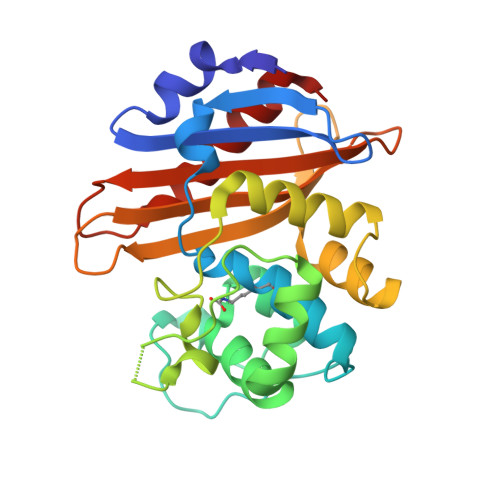
The 1.4 A crystal structure of the class D beta-lactamase OXA-1 complexed with doripenem.
Schneider, K.D., Karpen, M.E., Bonomo, R.A., Leonard, D.A., Powers, R.A.(2009) Biochemistry 48: 11840-11847
- PubMed: 19919101 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi901690r
- Primary Citation Related Structures:
3ISG - PubMed Abstract:
The clinical efficacy of carbapenem antibiotics depends on their resistance to the hydrolytic action of beta-lactamase enzymes. The structure of the class D beta-lactamase OXA-1 as an acyl complex with the carbapenem doripenem was determined to 1.4 A resolution. Unlike most class A and class C carbapenem complexes, the acyl carbonyl oxygen in the OXA-1-doripenem complex is bound in the oxyanion hole. Interestingly, no water molecules were observed in the vicinity of the acyl linkage, providing an explanation for why carbapenems inhibit OXA-1. The side chain amine of K70 remains fully carboxylated in the acyl structure, and the resulting carbamate group forms a hydrogen bond to the alcohol of the 6alpha-hydroxyethyl moiety of doripenem. The carboxylate attached to the beta-lactam ring of doripenem is stabilized by a salt bridge to K212 and a hydrogen bond with T213, in lieu of the interaction with an arginine side chain found in most other beta-lactamase-beta-lactam complexes (e.g., R244 in the class A member TEM-1). This novel set of interactions with the carboxylate results in a major shift of the carbapenem's pyrroline ring compared to the structure of the same ring in meropenem bound to OXA-13. Additionally, bond angles of the pyrroline ring suggest that after acylation, doripenem adopts the Delta(1) tautomer. These findings provide important insights into the role that carbapenems may have in the inactivation process of class D beta-lactamases.
- Department of Chemistry, Grand Valley State University, Allendale, Michigan 49401, USA.
Organizational Affiliation: