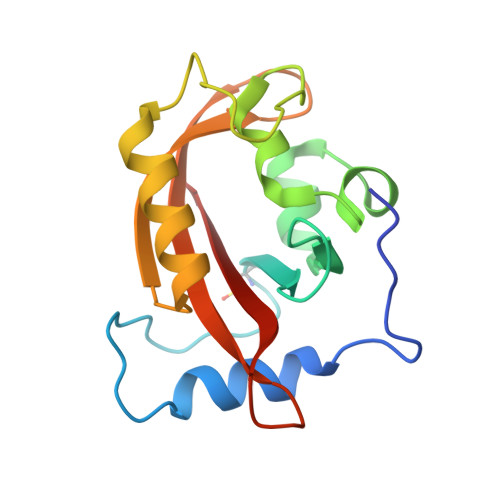
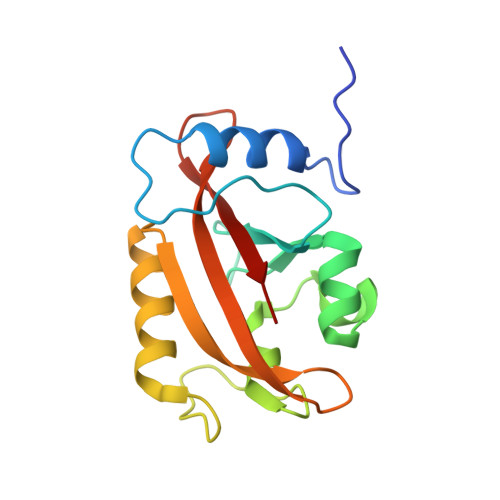
Illuminating solution responses of a LOV domain protein with photocoupled small-angle X-ray scattering.
Lamb, J.S., Zoltowski, B.D., Pabit, S.A., Li, L., Crane, B.R., Pollack, L.(2009) J Mol Biology 393: 909-919
- PubMed: 19712683 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.08.045
- Primary Citation Related Structures:
3IS2 - PubMed Abstract:
The PAS-LOV domain is a signal-transducing component found in a large variety of proteins that is responsible for sensing different stimuli such as light, oxygen, and voltage. The LOV protein VVD regulates blue light responses in the filamentous fungi Neurospora crassa. Using photocoupled, time-resolved small-angle X-ray scattering, we extract the solution protein structure in both dark-adapted and light-activated states. Two distinct dark-adapted conformations are detected in the wild-type protein: a compact structure that corresponds to the crystal structure of the dark-state monomer as well as an extended structure that is well modeled by introducing conformational disorder at the N-terminus of the protein. These conformations are accentuated in carefully selected variants, in which a key residue for propagating structural transitions, Cys71, has been mutated or oxidized. Despite different dark-state conformations, all proteins form a common dimer in response to illumination. Taken together, these data support a reaction scheme that describes the mechanism for light-induced dimerization of VVD. Envelope reconstructions of the transient light-state dimer reveal structures that are best described by a parallel arrangement of subunits that have significantly changed conformation compared to the crystal structure.
- School of Applied and Engineering Physics, Cornell University, Ithaca, NY 14853, USA.
Organizational Affiliation: