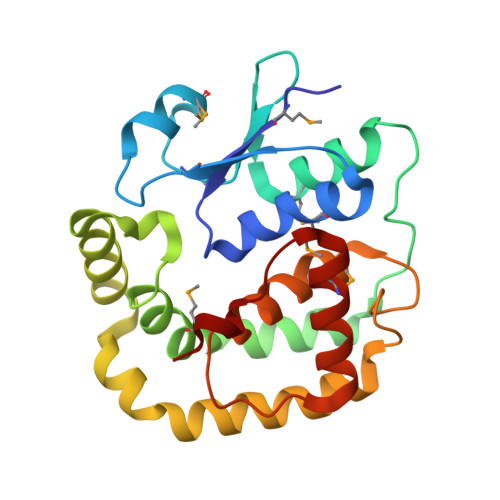
1.2 Angstrom Crystal Structure of the Glutaredoxin 2 (grxB) from Salmonella typhimurium in complex with Glutathione.
Minasov, G., Wawrzak, Z., Skarina, T., Onopriyenko, O., Peterson, S.N., Halavaty, A., Dauter, Z., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.