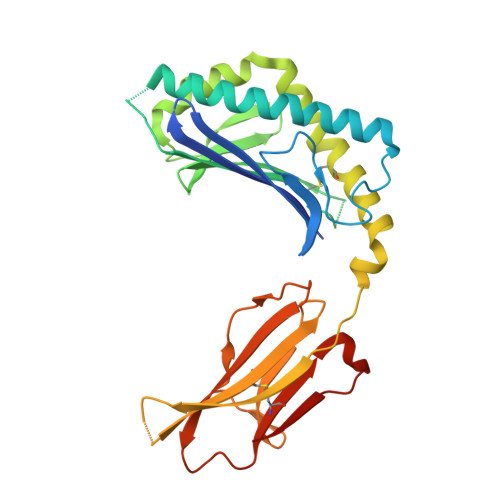
Lipid binding orientation within CD1d affects recognition of Borrelia burgorferi antigens by NKT cells.
Wang, J., Li, Y., Kinjo, Y., Mac, T.T., Gibson, D., Painter, G.F., Kronenberg, M., Zajonc, D.M.(2010) Proc Natl Acad Sci U S A 107: 1535-1540
- PubMed: 20080535 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0909479107
- Primary Citation Related Structures:
3ILP, 3ILQ - PubMed Abstract:
Invariant natural killer T cells (iNKT cells) respond to CD1d-presented glycolipids from Borrelia burgdorferi, the causative agent of Lyme disease. Although mouse and human iNKT cells respond to different antigens based on subtle differences in their fatty acids, the mechanism by which fatty acid structure determines antigenic potency is not well understood. Here we show that the mouse and human CD1d present glycolipids having different fatty acids, based in part upon a difference at a single amino acid position that is involved in positioning the sugar epitope. CD1d also can bind nonantigenic lipids, however, but unexpectedly, mouse CD1d orients the two aliphatic chains of a nonantigenic lipid rotated 180 degrees, causing a dramatic repositioning of the exposed sugar. Therefore, our data reveal the biochemical basis for the high degree of antigenic specificity of iNKT cells for certain fatty acids, and they suggest how microbes could alter fatty acid biosynthesis as an immune evasion mechanism.
- Division of Cell Biology, La Jolla Institute for Allergy and Immunology, La Jolla, CA 92037, USA.
Organizational Affiliation: