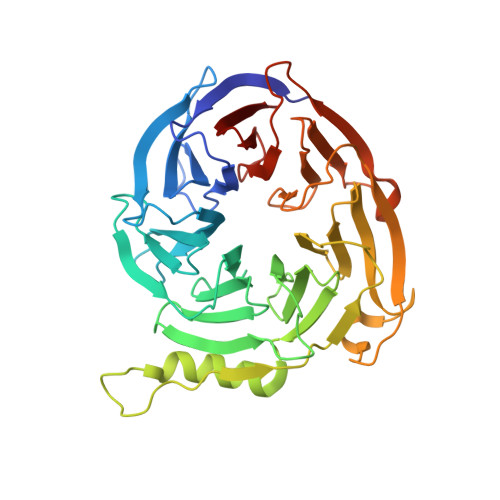
Role of the polycomb protein EED in the propagation of repressive histone marks.
Margueron, R., Justin, N., Ohno, K., Sharpe, M.L., Son, J., Drury, W.J., Voigt, P., Martin, S.R., Taylor, W.R., De Marco, V., Pirrotta, V., Reinberg, D., Gamblin, S.J.(2009) Nature 461: 762-767
- PubMed: 19767730 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature08398
- Primary Citation Related Structures:
3IIW, 3IIY, 3IJ0, 3IJ1, 3IJC - PubMed Abstract:
Polycomb group proteins have an essential role in the epigenetic maintenance of repressive chromatin states. The gene-silencing activity of the Polycomb repressive complex 2 (PRC2) depends on its ability to trimethylate lysine 27 of histone H3 (H3K27) by the catalytic SET domain of the EZH2 subunit, and at least two other subunits of the complex: SUZ12 and EED. Here we show that the carboxy-terminal domain of EED specifically binds to histone tails carrying trimethyl-lysine residues associated with repressive chromatin marks, and that this leads to the allosteric activation of the methyltransferase activity of PRC2. Mutations in EED that prevent it from recognizing repressive trimethyl-lysine marks abolish the activation of PRC2 in vitro and, in Drosophila, reduce global methylation and disrupt development. These findings suggest a model for the propagation of the H3K27me3 mark that accounts for the maintenance of repressive chromatin domains and for the transmission of a histone modification from mother to daughter cells.
- Howard Hughes Medical Institute and Department of Biochemistry, New York University Medical School, 522 First Avenue, New York, New York 10016, USA.
Organizational Affiliation: