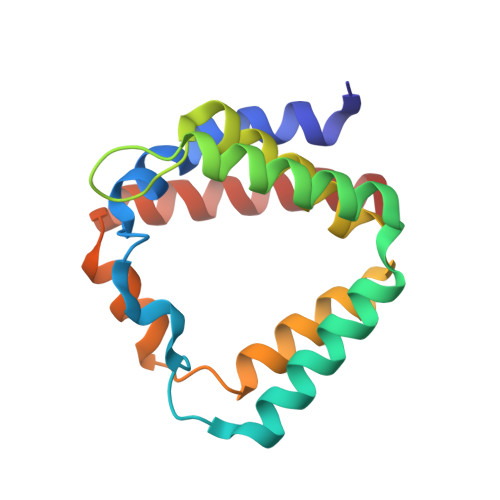
Identification of a single peridinin sensing Chl-a excitation in reconstituted PCP by crystallography and spectroscopy.
Schulte, T., Niedzwiedzki, D.M., Birge, R.R., Hiller, R.G., Polivka, T., Hofmann, E., Frank, H.A.(2009) Proc Natl Acad Sci U S A 106: 20764-20769
- PubMed: 19934052 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0908938106
- Primary Citation Related Structures:
3IIS, 3IIU - PubMed Abstract:
The peridinin-chlorophyll a-protein (PCP) of dinoflagellates is unique among the large variety of natural photosynthetic light-harvesting systems. In contrast to other chlorophyll protein complexes, the soluble PCP is located in the thylakoid lumen, and the carotenoid pigments outnumber the chlorophylls. The structure of the PCP complex consists of two symmetric domains, each with a central chlorophyll a (Chl-a) surrounded by four peridinin molecules. The protein provides distinctive surroundings for the pigment molecules, and in PCP, the specific environment around each peridinin results in overlapping spectral line shapes, suggestive of different functions within the protein. One particular Per, Per-614, is hypothesized to show the strongest electronic interaction with the central Chl-a. We have performed an in vitro reconstitution of pigments into recombinant PCP apo-protein (RFPCP) and into a mutated protein with an altered environment near Per-614. Steady-state and transient optical spectroscopic experiments comparing the RFPCP complex with the reconstituted mutant protein identify specific amino acid-induced spectral shifts. The spectroscopic assignments are reinforced by a determination of the structures of both RFPCP and the mutant by x-ray crystallography to a resolution better than 1.5 A. RFPCP and mutated RFPCP are unique in representing crystal structures of in vitro reconstituted light-harvesting pigment-protein complexes.
- Biophysics, Department of Biology and Biotechnology, Ruhr-University Bochum, D-44780 Bochum, Germany.
Organizational Affiliation: