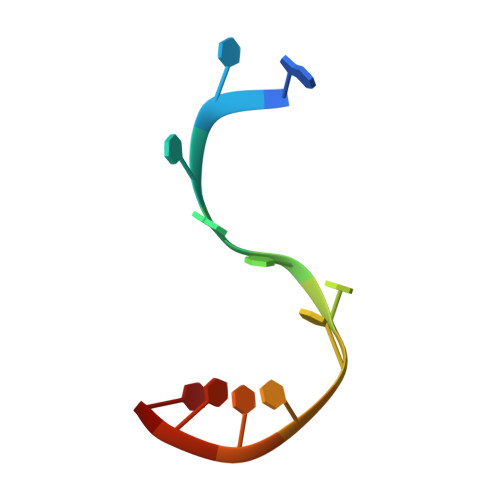
Structure of ERA in complex with the 3' end of 16S rRNA: implications for ribosome biogenesis
Tu, C., Zhou, X., Tropea, J.E., Austin, B.P., Waugh, D.S., Court, D.L., Ji, X.(2009) Proc Natl Acad Sci U S A 106: 14843-14848
- PubMed: 19706445 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0904032106
- Primary Citation Related Structures:
3IEU, 3IEV - PubMed Abstract:
ERA, composed of an N-terminal GTPase domain followed by an RNA-binding KH domain, is essential for bacterial cell viability. It binds to 16S rRNA and the 30S ribosomal subunit. However, its RNA-binding site, the functional relationship between the two domains, and its role in ribosome biogenesis remain unclear. We have determined two crystal structures of ERA, a binary complex with GDP and a ternary complex with a GTP-analog and the 1531AUCACCUCCUUA1542 sequence at the 3' end of 16S rRNA. In the ternary complex, the first nine of the 12 nucleotides are recognized by the protein. We show that GTP binding is a prerequisite for RNA recognition by ERA and that RNA recognition stimulates its GTP-hydrolyzing activity. Based on these and other data, we propose a functional cycle of ERA, suggesting that the protein serves as a chaperone for processing and maturation of 16S rRNA and a checkpoint for assembly of the 30S ribosomal subunit. The AUCA sequence is highly conserved among bacteria, archaea, and eukaryotes, whereas the CCUCC, known as the anti-Shine-Dalgarno sequence, is conserved in noneukaryotes only. Therefore, these data suggest a common mechanism for a highly conserved ERA function in all three kingdoms of life by recognizing the AUCA, with a "twist" for noneukaryotic ERA proteins by also recognizing the CCUCC.
- Center for Cancer Research, National Cancer Institute, National Institutes of Health, Frederick, MD 21702, USA.
Organizational Affiliation: