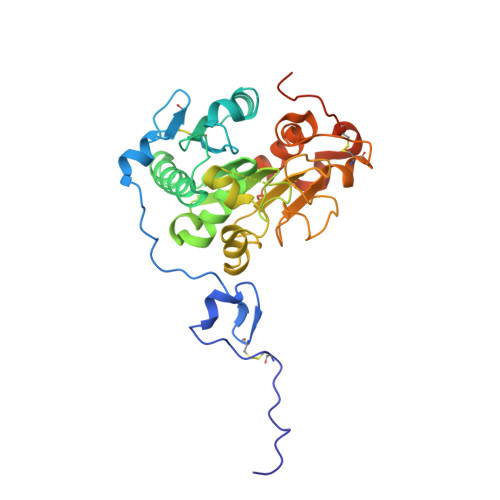
Structural redesign of lipase B from Candida antarctica by circular permutation and incremental truncation.
Qian, Z., Horton, J.R., Cheng, X., Lutz, S.(2009) J Mol Biology 393: 191-201
- PubMed: 19683009 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.08.008
- Primary Citation Related Structures:
3ICV, 3ICW - PubMed Abstract:
Circular permutation of Candida antarctica lipase B yields several enzyme variants with substantially increased catalytic activity. To better understand the structural and functional consequences of protein termini reorganization, we have applied protein engineering and x-ray crystallography to cp283, one of the most active hydrolase variants. Our initial investigation has focused on the role of an extended surface loop, created by linking the native N- and C-termini, on protein integrity. Incremental truncation of the loop partially compensates for observed losses in secondary structure and the permutants' temperature of unfolding. Unexpectedly, the improvements are accompanied by quaternary-structure changes from monomer to dimer. The crystal structures of one truncated variant (cp283 Delta 7) in the apo-form determined at 1.49 A resolution and with a bound phosphonate inhibitor at 1.69 A resolution confirmed the formation of a homodimer by swapping of the enzyme's 35-residue N-terminal region. Separately, the new protein termini at amino acid positions 282/283 convert the narrow access tunnel to the catalytic triad into a broad crevice for accelerated substrate entry and product exit while preserving the native active-site topology for optimal catalytic turnover.
- Department of Chemistry, Emory University, 1515 Dickey Drive, Atlanta, GA 30322, USA.
Organizational Affiliation: