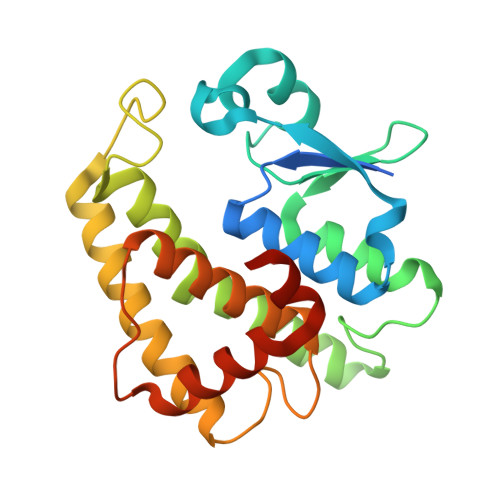
Structures of yeast glutathione-S-transferase Gtt2 reveal a new catalytic type of GST family.
Ma, X.X., Jiang, Y.L., He, Y.X., Bao, R., Chen, Y.X., Zhou, C.Z.(2009) EMBO Rep 10: 1320-1326
- PubMed: 19851333 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2009.216
- Primary Citation Related Structures:
3ERF, 3ERG, 3IBH - PubMed Abstract:
Glutathione-S-transferases (GSTs) are ubiquitous detoxification enzymes that catalyse the conjugation of electrophilic substrates to glutathione. Here, we present the crystal structures of Gtt2, a GST of Saccharomyces cerevisiae, in apo and two ligand-bound forms, at 2.23 A, 2.20 A and 2.10 A, respectively. Although Gtt2 has the overall structure of a GST, the absence of the classic catalytic essential residues--tyrosine, serine and cysteine--distinguishes it from all other cytosolic GSTs of known structure. Site-directed mutagenesis in combination with activity assays showed that instead of the classic catalytic residues, a water molecule stabilized by Ser129 and His123 acts as the deprotonator of the glutathione sulphur atom. Furthermore, only glycine and alanine are allowed at the amino-terminus of helix-alpha1 because of stereo-hindrance. Taken together, these results show that yeast Gtt2 is a novel atypical type of cytosolic GST.
- Hefei National Laboratory for Physical Sciences at Microscale, and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230027, People's Republic of China.
Organizational Affiliation: