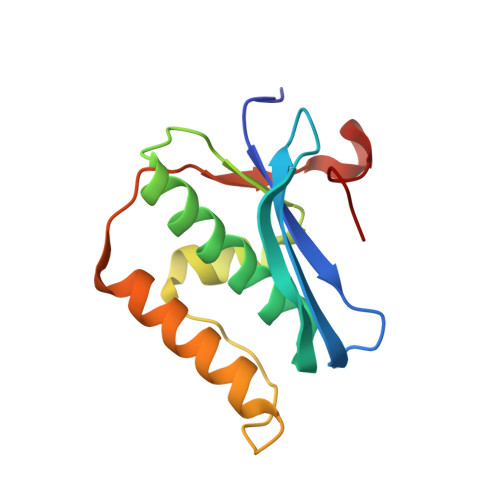
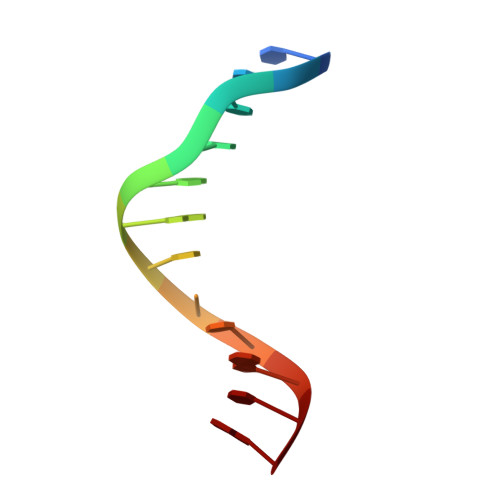
Pairing geometry of the hydrophobic thymine analogue 2,4-difluorotoluene in duplex DNA as analyzed by X-ray crystallography.
Pallan, P.S., Egli, M.(2009) J Am Chem Soc 131: 12548-12549
- PubMed: 19685868 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja905739j
- Primary Citation Related Structures:
3I8D - PubMed Abstract:
Certain DNA polymerases (pols) were found to efficiently insert A opposite the hydrophobic T isostere 2,4-difluorotoluene (F) and vice versa, resulting in the widely held belief that some pols rely on shape rather than H-bonding for accurate replication. Using X-ray crystallography we have analyzed the geometry of F:A pairs in duplex DNA and observed a distance between fluorine and the exocyclic amino group of A that is consistent with a H-bond, thus challenging the assumption that the F analogue is unable to engage in H-bonding as well as the steric hypothesis of DNA replication. Therefore, shape and H-bonding are inherently related, and steric constraints at a pol active site, or conferred by stacking or the DNA backbone conformation, may enable H-bonding by F.
- Department of Biochemistry, School of Medicine, Vanderbilt University, Nashville, Tennessee 37232, USA.
Organizational Affiliation: