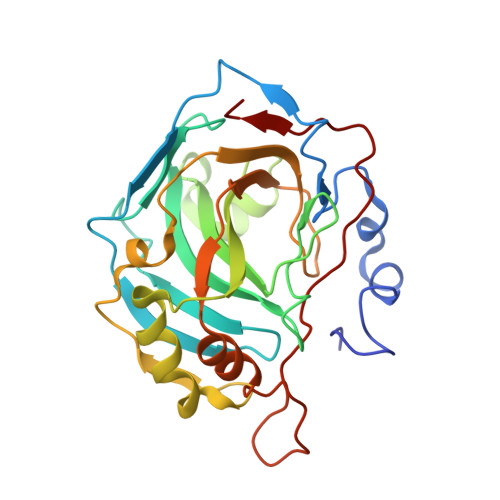
High-resolution structure of human carbonic anhydrase II complexed with acetazolamide reveals insights into inhibitor drug design.
Sippel, K.H., Robbins, A.H., Domsic, J., Genis, C., Agbandje-McKenna, M., McKenna, R.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 992-995
- PubMed: 19851004 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109036665
- Primary Citation Related Structures:
3HS4 - PubMed Abstract:
The crystal structure of human carbonic anhydrase II (CA II) complexed with the inhibitor acetazolamide (AZM) has been determined at 1.1 A resolution and refined to an R(cryst) of 11.2% and an R(free) of 14.7%. As observed in previous CA II-inhibitor complexes, AZM binds directly to the zinc and makes several key interactions with active-site residues. The high-resolution data also showed a glycerol molecule adjacent to the AZM in the active site and two additional AZMs that are adventitiously bound on the surface of the enzyme. The co-binding of AZM and glycerol in the active site demonstrate that given an appropriate ring orientation and substituents, an isozyme-specific CA inhibitor may be developed.
- Department of Biochemistry and Molecular Biology, College of Medicine, University of Florida, Gainesville, FL 32610, USA.
Organizational Affiliation: