The hunt for 8-oxoguanine deaminase.
Hall, R.S., Fedorov, A.A., Marti-Arbona, R., Fedorov, E.V., Kolb, P., Sauder, J.M., Burley, S.K., Shoichet, B.K., Almo, S.C., Raushel, F.M.(2010) J Am Chem Soc 132: 1762-1763
- PubMed: 20088583 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja909817d
- Primary Citation Related Structures:
3HPA - PubMed Abstract:
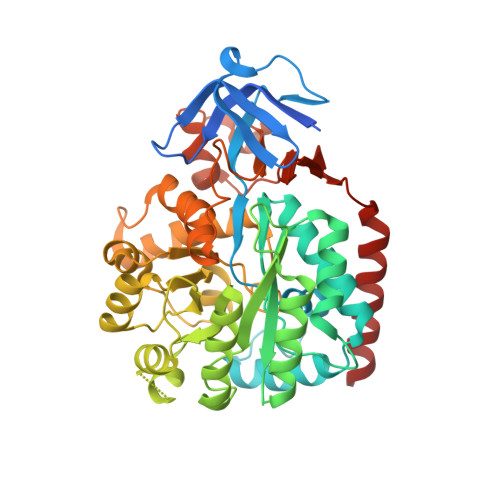
An enzyme from Pseudomonas aeruginosa, Pa0142 (gi|9945972), that is able to catalyze the deamination of 8-oxoguanine (8-oxoG) to uric acid has been identified for the first time. 8-Oxoguanine is formed by the oxidation of guanine residues within DNA by reactive oxygen species, and this lesion results in G:C to T:A transversions. The value of k(cat)/K(m) for the deamination of 8-oxoG by Pa0142 at pH 8.0 and 30 degrees C is 2.0 x 10(4) M(-1) s(-1). This enzyme can also catalyze the deamination of isocystosine and guanine at rates that are approximately an order of magnitude lower. The three-dimensional structure of a homologous enzyme (gi|44264246) from the Sargasso Sea has been determined by X-ray diffraction methods to a resolution of 2.2 A (PDB entry). The enzyme folds as a (beta/alpha)(8) barrel and is a member of the amidohydrolase superfamily with a single zinc in the active site. This enzyme catalyzes the deamination of 8-oxoG with a k(cat)/K(m) value of 2.7 x 10(5) M(-1) s(-1). Computational docking of potential high-energy intermediates for the deamination reaction to the X-ray crystal structure suggests that active-site binding of 8-oxoG is facilitated by hydrogen-bond interactions from a conserved glutamine that follows beta-strand 1 with the carbonyl group at C6, a conserved tyrosine that follows beta-strand 2 with N7, and a conserved cysteine residue that follows beta-strand 4 with the carbonyl group at C8. A bioinformatic analysis of available protein sequences suggests that approximately 200 other bacteria possess an enzyme capable of catalyzing the deamination of 8-oxoG.
- Department of Chemistry, Texas A&M University, P.O. Box 30012, College Station, Texas 77842-3012, USA.
Organizational Affiliation: