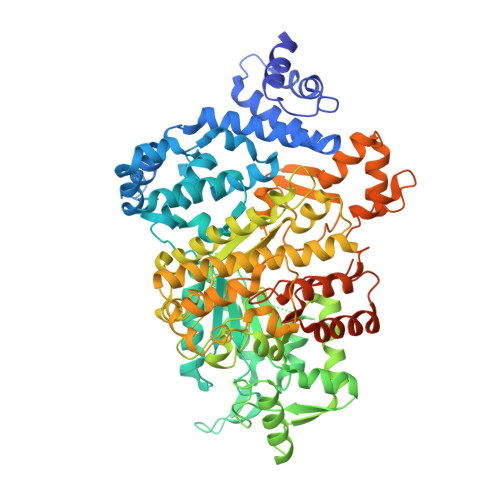
Structural basis for allosteric regulation of human ribonucleotide reductase by nucleotide-induced oligomerization.
Fairman, J.W., Wijerathna, S.R., Ahmad, M.F., Xu, H., Nakano, R., Jha, S., Prendergast, J., Welin, R.M., Flodin, S., Roos, A., Nordlund, P., Li, Z., Walz, T., Dealwis, C.G.(2011) Nat Struct Mol Biol 18: 316-322
- PubMed: 21336276 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2007
- Primary Citation Related Structures:
2WGH, 3HNC, 3HND, 3HNE, 3HNF, 3PAW - PubMed Abstract:
Ribonucleotide reductase (RR) is an α(n)β(n) (RR1-RR2) complex that maintains balanced dNTP pools by reducing NDPs to dNDPs. RR1 is the catalytic subunit, and RR2 houses the free radical required for catalysis. RR is allosterically regulated by its activator ATP and its inhibitor dATP, which regulate RR activity by inducing oligomerization of RR1. Here, we report the first X-ray structures of human RR1 bound to TTP alone, dATP alone, TTP-GDP, TTP-ATP, and TTP-dATP. These structures provide insights into regulation of RR by ATP or dATP. At physiological dATP concentrations, RR1 forms inactive hexamers. We determined the first X-ray structure of the RR1-dATP hexamer and used single-particle electron microscopy to visualize the α(6)-ββ'-dATP holocomplex. Site-directed mutagenesis and functional assays confirm that hexamerization is a prerequisite for inhibition by dATP. Our data indicate a mechanism for regulating RR activity by dATP-induced oligomerization.
- Department of Biochemistry, University of Tennessee, Knoxville, TN, USA.
Organizational Affiliation: