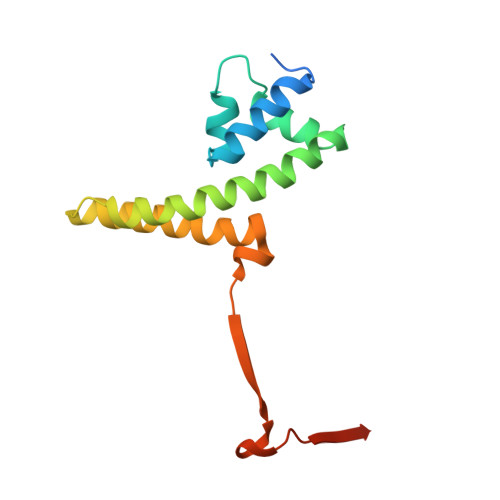
Crystal structure of the DNA-recognition component of the bacterial virus Sf6 genome-packaging machine
Zhao, H., Finch, C.J., Sequeira, R.D., Johnson, B.A., Johnson, J.E., Casjens, S.R., Tang, L.(2010) Proc Natl Acad Sci U S A 107: 1971-1976
- PubMed: 20133842 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0908569107
- Primary Citation Related Structures:
3HEF - PubMed Abstract:
In herpesviruses and many bacterial viruses, genome-packaging is a precisely mediated process fulfilled by a virally encoded molecular machine called terminase that consists of two protein components: A DNA-recognition component that defines the specificity for packaged DNA, and a catalytic component that provides energy for the packaging reaction by hydrolyzing ATP. The terminase docks onto the portal protein complex embedded in a single vertex of a preformed viral protein shell called procapsid, and pumps the viral DNA into the procapsid through a conduit formed by the portal. Here we report the 1.65 A resolution structure of the DNA-recognition component gp1 of the Shigella bacteriophage Sf6 genome-packaging machine. The structure reveals a ring-like octamer formed by interweaved protein monomers with a highly extended fold, embracing a tunnel through which DNA may be translocated. The N-terminal DNA-binding domains form the peripheral appendages surrounding the octamer. The central domain contributes to oligomerization through interactions of bundled helices. The C-terminal domain forms a barrel with parallel beta-strands. The structure reveals a common scheme for oligomerization of terminase DNA-recognition components, and provides insights into the role of gp1 in formation of the packaging-competent terminase complex and assembly of the terminase with the portal, in which ring-like protein oligomers stack together to form a continuous channel for viral DNA translocation.
- Department of Molecular Biosciences, University of Kansas, 1200 Sunnyside Avenue, Lawrence, KS 66045, USA.
Organizational Affiliation: