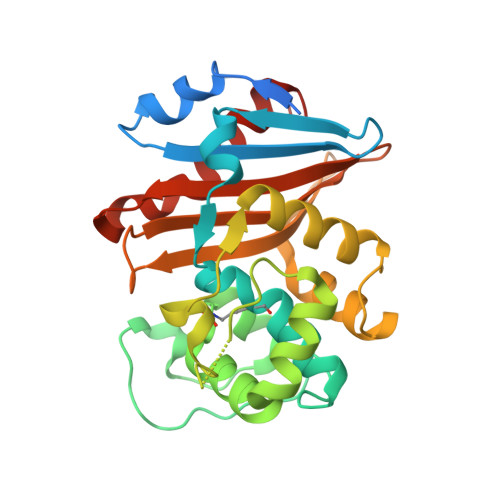
Crystal structure of the OXA-48 beta-lactamase reveals mechanistic diversity among class D carbapenemases.
Docquier, J.D., Calderone, V., De Luca, F., Benvenuti, M., Giuliani, F., Bellucci, L., Tafi, A., Nordmann, P., Botta, M., Rossolini, G.M., Mangani, S.(2009) Chem Biol 16: 540-547
- PubMed: 19477418 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2009.04.010
- Primary Citation Related Structures:
3HBR - PubMed Abstract:
Carbapenem-hydrolyzing class D beta-lactamases (CHDLs) are enzymes found in important Gram-negative pathogens (mainly Acinetobacter baumannii and Enterobacteriaceae) that confer resistance to beta-lactam antibiotics, and notably carbapenems. The crystal structure of the OXA-48 carbapenemase was determined at pH 7.5 and at a resolution of 1.9 A. Surprisingly, and by contrast with OXA-24, the only other CHDL of known crystal structure, the structure of OXA-48 was similar to OXA-10, an enzyme devoid of carbapenemase activity, indicating that the hydrolysis of these compounds could depend on subtle changes in the active site region. Moreover, the active site groove of OXA-48 was different from that of OXA-24 in shape, dimensions, and charge distribution. Molecular dynamics pointed to the functional relevance of residues located in or close to the beta5-beta6 loop and allowed us to propose a mechanism for carbapenem hydrolysis by OXA-48.
- Dipartimento di Biologia Molecolare, Università di Siena, Italy.
Organizational Affiliation: