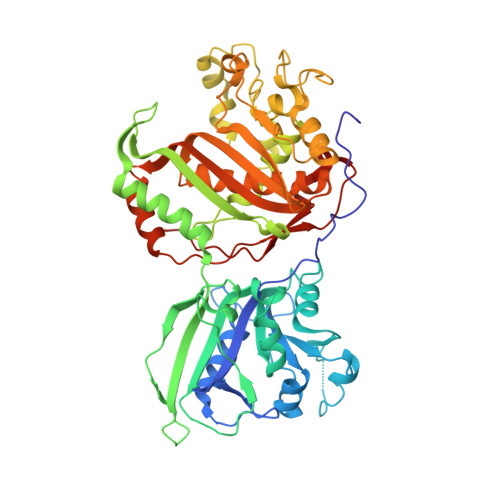
Structures of dihydrofolate reductase-thymidylate synthase of Trypanosoma cruzi in the folate-free state and in complex with two antifolate drugs, trimetrexate and methotrexate.
Senkovich, O., Schormann, N., Chattopadhyay, D.(2009) Acta Crystallogr D Biol Crystallogr 65: 704-716
- PubMed: 19564691 Search on PubMed
- DOI: https://doi.org/10.1107/S090744490901230X
- Primary Citation Related Structures:
3HBB - PubMed Abstract:
The flagellate protozoan parasite Trypanosoma cruzi is the pathogenic agent of Chagas disease (also called American trypanosomiasis), which causes approximately 50,000 deaths annually. The disease is endemic in South and Central America. The parasite is usually transmitted by a blood-feeding insect vector, but can also be transmitted via blood transfusion. In the chronic form, Chagas disease causes severe damage to the heart and other organs. There is no satisfactory treatment for chronic Chagas disease and no vaccine is available. There is an urgent need for the development of chemotherapeutic agents for the treatment of T. cruzi infection and therefore for the identification of potential drug targets. The dihydrofolate reductase activity of T. cruzi, which is expressed as part of a bifunctional enzyme, dihydrofolate reductase-thymidylate synthase (DHFR-TS), is a potential target for drug development. In order to gain a detailed understanding of the structure-function relationship of T. cruzi DHFR, the three-dimensional structure of this protein in complex with various ligands is being studied. Here, the crystal structures of T. cruzi DHFR-TS with three different compositions of the DHFR domain are reported: the folate-free state, the complex with the lipophilic antifolate trimetrexate (TMQ) and the complex with the classical antifolate methotrexate (MTX). These structures reveal that the enzyme is a homodimer with substantial interactions between the two TS domains of neighboring subunits. In contrast to the enzymes from Cryptosporidium hominis and Plasmodium falciparum, the DHFR and TS active sites of T. cruzi lie on the same side of the monomer. As in other parasitic DHFR-TS proteins, the N-terminal extension of the T. cruzi enzyme is involved in extensive interactions between the two domains. The DHFR active site of the T. cruzi enzyme shows subtle differences compared with its human counterpart. These differences may be exploited for the development of antifolate-based therapeutic agents for the treatment of T. cruzi infection.
- Center for Biophysical Sciences and Engineering, University of Alabama at Birmingham, Birmingham, AL 35294, USA.
Organizational Affiliation: