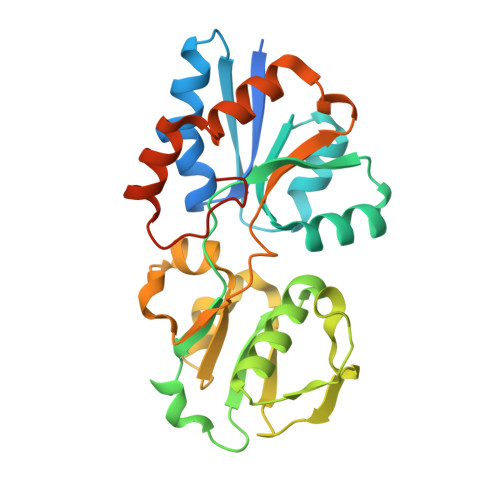
Crystal structure of lipoprotein GNA1946 from Neisseria meningitidis
Yang, X., Wu, Z., Wang, X., Yang, C., Xu, H., Shen, Y.(2009) J Struct Biol 168: 437-443
- PubMed: 19733245 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2009.09.001
- Primary Citation Related Structures:
3GXA, 3IR1 - PubMed Abstract:
GNA1946, a conserved outer membrane lipoprotein from Neisseria meningitidis, has been identified as a candidate antigen for an urgently needed broad-spectrum meningococcal vaccine. It has been predicted to be a periplasmic receptor in the D-methionine uptake ABC transporter system. The crystal structure of GNA1946 was solved by the single-wavelength anomalous dispersion (SAD) method to a resolution of 2.25 A, and it reveals a Venus flytrap-like structure. GNA1946 consists of two globular lobes connected by a hinge region. Surprisingly, the structure showed an L-methionine bound within the cleft between the lobes. A comparison of GNA1946 with two other outer membrane lipoproteins, the L-methionine-binding Tp32 from Treponema pallidum and the dipeptide GlyMet-binding protein Pg110 from Staphylococcus aureus, revealed that although these three proteins share low sequence similarities, there is a high degree of structural conservation and similar substrate-binding frameworks. Our results reveal that GNA1946 is an L-methionine binding lipoprotein in the outer membrane, and should function as an initial receptor for ABC transporters with high affinity and specificity. The GNA1946 structure reported here should provide a valuable starting point for the development of a broad-spectrum meningococcal vaccine.
- Tianjin Key Laboratory of Protein Science, The College of Life Science, Nankai University, Tianjin, China.
Organizational Affiliation: