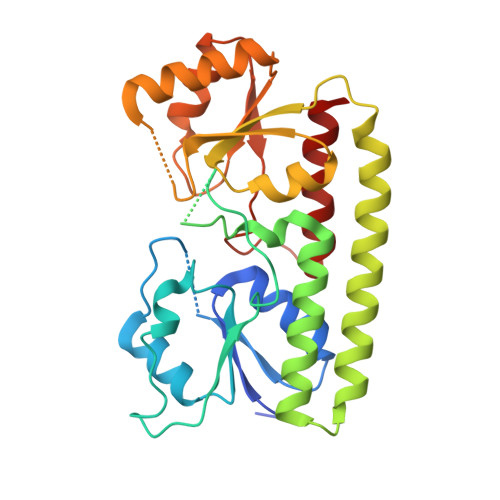
The laminin-binding protein Lbp from Streptococcus pyogenes is a zinc receptor
Linke, C., Caradoc-Davies, T.T., Young, P.G., Proft, T., Baker, E.N.(2009) J Bacteriol 191: 5814-5823
- PubMed: 19617361 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00485-09
- Primary Citation Related Structures:
3GI1 - PubMed Abstract:
The common pathogen Streptococcus pyogenes colonizes the human skin and tonsils and can invade underlying tissues. This requires the adhesion of S. pyogenes to host surface receptors mediated through adhesins. The laminin-binding protein Lbp has been suggested as an adhesin, specific for the human extracellular matrix protein laminin. Sequence alignments, however, indicate a relationship between Lbp and a family of bacterial metal-binding receptors. To further analyze the role of Lbp in S. pyogenes and its potential role in pathogenicity, Lbp has been crystallized, and its structure has been solved at a resolution of 2.45 A (R = 0.186; R(free) = 0.251). Lbp has the typical metal-binding receptor fold, comprising two globular (beta/alpha)(4) domains connected by a helical backbone. The two domains enclose the metal-binding site, which contains a zinc ion. The interaction of Lbp with laminin was further investigated and shown to be specific in vitro. Localization studies with antibodies specific for Lbp show that the protein is attached to the membrane. The data suggest that Lbp is primarily a zinc-binding protein, and we suggest that its interaction with laminin in vivo may be mediated via zinc bound to laminin.
- School of Biological Sciences, University of Auckland, Private Bag 92019, Auckland, New Zealand.
Organizational Affiliation: