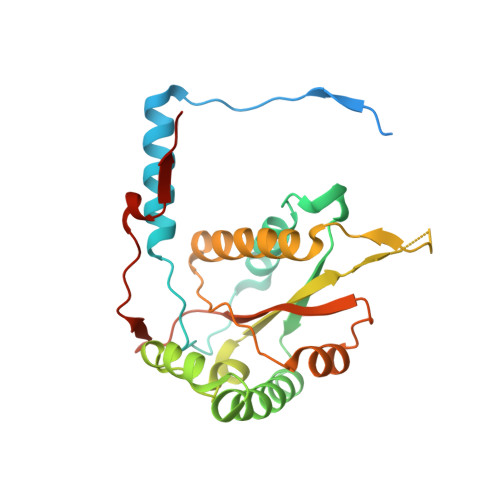
Crystal structure of iodotyrosine deiodinase, a novel flavoprotein responsible for iodide salvage in thyroid glands.
Thomas, S.R., McTamney, P.M., Adler, J.M., Laronde-Leblanc, N., Rokita, S.E.(2009) J Biological Chem 284: 19659-19667
- PubMed: 19436071 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.013458
- Primary Citation Related Structures:
3GB5, 3GFD, 3GH8 - PubMed Abstract:
The flavoprotein iodotyrosine deiodinase (IYD) salvages iodide from mono- and diiodotyrosine formed during the biosynthesis of the thyroid hormone thyroxine. Expression of a soluble domain of this membrane-bound enzyme provided sufficient material for crystallization and characterization by x-ray diffraction. The structures of IYD and two co-crystals containing substrates, mono- and diiodotyrosine, alternatively, were solved at resolutions of 2.0, 2.45, and 2.6 A, respectively. The structure of IYD is homologous to others in the NADH oxidase/flavin reductase superfamily, but the position of the active site lid in IYD defines a new subfamily within this group that includes BluB, an enzyme associated with vitamin B(12) biosynthesis. IYD and BluB also share key interactions involving their bound flavin mononucleotide that suggest a unique catalytic behavior within the superfamily. Substrate coordination to IYD induces formation of an additional helix and coil that act as an active site lid to shield the resulting substrate.flavin complex from solvent. This complex is stabilized by aromatic stacking and extensive hydrogen bonding between the substrate and flavin. The carbon-iodine bond of the substrate is positioned directly over the C-4a/N-5 region of the flavin to promote electron transfer. These structures now also provide a molecular basis for understanding thyroid disease based on mutations of IYD.
- Department of Chemistry and Biochemistry, Center for Biomolecular Structure and Organization, University of Maryland, College Park, Maryland 20742, USA.
Organizational Affiliation: