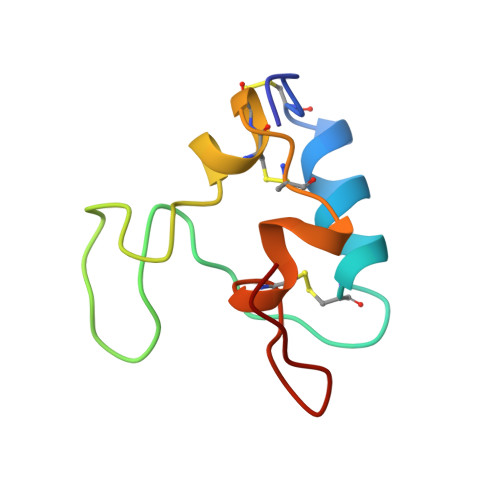
Solution structure of human insulin-like growth factor 1: a nuclear magnetic resonance and restrained molecular dynamics study.
Cooke, R.M., Harvey, T.S., Campbell, I.D.(1991) Biochemistry 30: 5484-5491
- PubMed: 2036417 Search on PubMed
- DOI: https://doi.org/10.1021/bi00236a022
- Primary Citation Related Structures:
2GF1, 3GF1 - PubMed Abstract:
The solution structure of human insulin-like growth factor 1 has been investigated with a combination of nuclear magnetic resonance and restrained molecular dynamics methods. The results show that the solution structure is similar to that of insulin, but minor differences exist. The regions homologous to insulin are well-defined, while the remainder of the molecule exhibits greater disorder. The resultant structures have been used to visualize the sites for interaction with a number of physiologically important proteins.
- Department of Biochemistry, University of Oxford, U.K.
Organizational Affiliation: