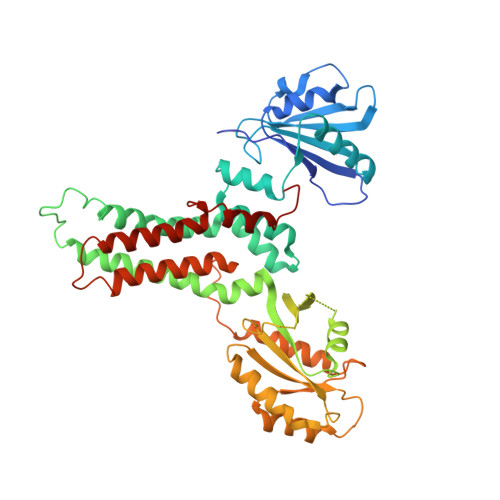
Kissing G domains of MnmE monitored by X-ray crystallography and pulse electron paramagnetic resonance spectroscopy
Meyer, S., Bohme, S., Kruger, A., Steinhoff, H.-J., Klare, J.P., Wittinghofer, A.(2009) PLoS Biol 7: e1000212-e1000212
- PubMed: 19806182 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1000212
- Primary Citation Related Structures:
3GEE, 3GEH, 3GEI - PubMed Abstract:
MnmE, which is involved in the modification of the wobble position of certain tRNAs, belongs to the expanding class of G proteins activated by nucleotide-dependent dimerization (GADs). Previous models suggested the protein to be a multidomain protein whose G domains contact each other in a nucleotide dependent manner. Here we employ a combined approach of X-ray crystallography and pulse electron paramagnetic resonance (EPR) spectroscopy to show that large domain movements are coupled to the G protein cycle of MnmE. The X-ray structures show MnmE to be a constitutive homodimer where the highly mobile G domains face each other in various orientations but are not in close contact as suggested by the GDP-AlF(x) structure of the isolated domains. Distance measurements by pulse double electron-electron resonance (DEER) spectroscopy show that the G domains adopt an open conformation in the nucleotide free/GDP-bound and an open/closed two-state equilibrium in the GTP-bound state, with maximal distance variations of 18 A. With GDP and AlF(x), which mimic the transition state of the phosphoryl transfer reaction, only the closed conformation is observed. Dimerization of the active sites with GDP-AlF(x) requires the presence of specific monovalent cations, thus reflecting the requirements for the GTPase reaction of MnmE. Our results directly demonstrate the nature of the conformational changes MnmE was previously suggested to undergo during its GTPase cycle. They show the nucleotide-dependent dynamic movements of the G domains around two swivel positions relative to the rest of the protein, and they are of crucial importance for understanding the mechanistic principles of this GAD.
- Department of Structural Biology, Max-Planck-Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: