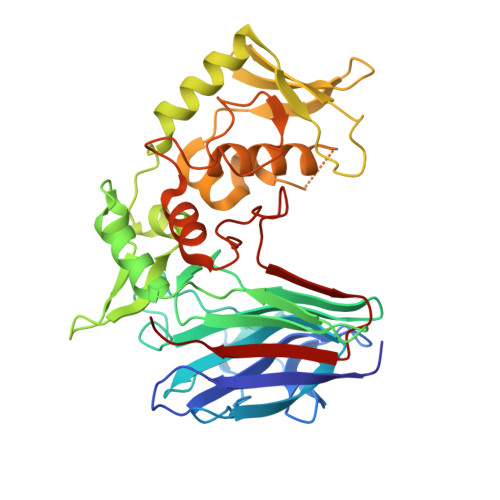
Structure, mechanistic action, and essential residues of a GH-64 enzyme, laminaripentaose-producing beta-1,3-glucanase.
Wu, H.M., Liu, S.W., Hsu, M.T., Hung, C.L., Lai, C.C., Cheng, W.C., Wang, H.J., Li, Y.K., Wang, W.C.(2009) J Biological Chem 284: 26708-26715
- PubMed: 19640850 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.010983
- Primary Citation Related Structures:
3GD0, 3GD9 - PubMed Abstract:
Laminaripentaose-producing beta-1,3-glucanase (LPHase), a member of glycoside hydrolase family 64, cleaves a long-chain polysaccharide beta-1,3-glucan into specific pentasaccharide oligomers. The crystal structure of LPHase from Streptomyces matensis DIC-108 was solved to 1.62 A resolution using multiple-wavelength anomalous dispersion methods. The LPHase structure reveals a novel crescent-like fold; it consists of a barrel domain and a mixed (alpha/beta) domain, forming a wide-open groove between the two domains. The liganded crystal structure was also solved to 1.80 A, showing limited conformational changes. Within the wide groove, a laminaritetraose molecule is found to sit in an electronegatively charged central region and is proximal to several conserved residues including two carboxylates (Glu(154) and Asp(170)) and four other sugar-binding residues (Thr(156), Asn(158), Trp(163), and Thr(167)). Molecular modeling using a laminarihexaose as a substrate suggests roles for Glu(154) and Asp(170) as acid and base catalysts, respectively, whereas the side chains of Thr(156), Asn(158), and Trp(163) demarcate subsite +5. Site-directed mutagenesis of Glu(154) and Asp(170) confirms that both carboxylates are essential for catalysis. Together, our results suggest that LPHase uses a direct displacement mechanism involving Glu(154) and Asp(170) to cleave a beta-1,3-glucan into specific alpha-pentasaccharide oligomers.
- Department of Life Science, Institute of Molecular and Cellular Biology, National Tsing Hua University, Hsinchu 300, Taiwan.
Organizational Affiliation: