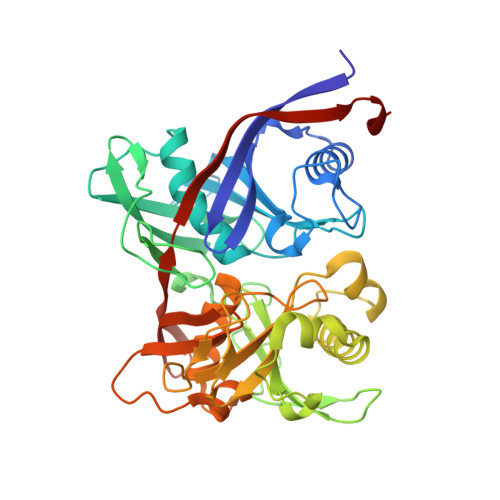
Crystal structure and putative mechanism of 3-methylitaconate-delta-isomerase from Eubacterium barkeri
Velarde, M., Macieira, S., Hilberg, M., Broker, G., Tu, S.-M., Golding, B.T., Pierik, A.J., Buckel, W., Messerschmidt, A.(2009) J Mol Biology 391: 609-620
- PubMed: 19559030 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.06.052
- Primary Citation Related Structures:
3G7K - PubMed Abstract:
3-Methylitaconate-Delta-isomerase (Mii) participates in the nicotinate fermentation pathway of the anaerobic soil bacterium Eubacterium barkeri (order Clostridiales) by catalyzing the reversible conversion of (R)-3-methylitaconate (2-methylene-3-methylsuccinate) to 2,3-dimethylmaleate. The enzyme is also able to catalyze the isomerization of itaconate (methylenesuccinate) to citraconate (methylmaleate) with ca 10-fold higher K(m) but > 1000-fold lower k(cat). The gene mii from E. barkeri was cloned and expressed in Escherichia coli. The protein produced with a C-terminal Strep-tag exhibited the same specific activity as the wild-type enzyme. The crystal structure of Mii from E. barkeri has been solved at a resolution of 2.70 A. The asymmetric unit of the P2(1)2(1)2(1) unit cell with parameters a = 53.1 A, b = 142.3 A, and c = 228.4 A contains four molecules of Mii. The enzyme belongs to a group of isomerases with a common structural feature, the so-called diaminopimelate epimerase fold. The monomer of 380 amino acid residues has two topologically similar domains exhibiting an alpha/beta-fold. The active site is situated in a cleft between these domains. The four Mii molecules are arranged as a tetramer with 222 symmetry for the N-terminal domains. The C-terminal domains have different relative positions with respect to the N-terminal domains resulting in a closed conformation for molecule A and two distinct open conformations for molecules B and D. The C-terminal domain of molecule C is disordered. The Mii active site contains the putative catalytic residues Lys62 and Cys96, for which mechanistic roles are proposed based on a docking experiment of the Mii substrate complex. The active sites of Mii and the closely related PrpF, most likely a methylaconitate Delta-isomerase, have been compared. The overall architecture including the active-site Lys62, Cys96, His300, and Ser17 (Mii numbering) is similar. This positioning of (R)-3-methylitaconate allows Cys96 (as thiolate) to deprotonate C-3 and (as thiol) to donate a proton to the methylene carbon atom of the resulting allylic carbanion. Interestingly, the active site of isopentenyl diphosphate isomerase type I also contains a cysteine that cooperates with glutamate rather than lysine. It has been proposed that the initial step in this enzyme is a protonation generating a tertiary carbocation intermediate.
- Department of Proteomics and Signal Transduction, Max-Planck-Institute of Biochemistry, 82152 Martinsried, Germany.
Organizational Affiliation: