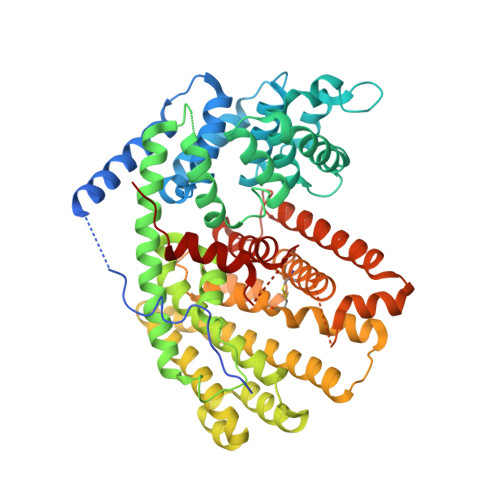
Crystal structure of (+)-delta-cadinene synthase from Gossypium arboreum and evolutionary divergence of metal binding motifs for catalysis.
Gennadios, H.A., Gonzalez, V., Di Costanzo, L., Li, A., Yu, F., Miller, D.J., Allemann, R.K., Christianson, D.W.(2009) Biochemistry 48: 6175-6183
- PubMed: 19489610 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi900483b
- Primary Citation Related Structures:
3G4D, 3G4F - PubMed Abstract:
(+)-Delta-cadinene synthase (DCS) from Gossypium arboreum (tree cotton) is a sesquiterpene cyclase that catalyzes the cyclization of farnesyl diphosphate in the first committed step of the biosynthesis of gossypol, a phytoalexin that defends the plant from bacterial and fungal pathogens. Here, we report the X-ray crystal structure of unliganded DCS at 2.4 A resolution and the structure of its complex with three putative Mg(2+) ions and the substrate analogue inhibitor 2-fluorofarnesyl diphosphate (2F-FPP) at 2.75 A resolution. These structures illuminate unusual features that accommodate the trinuclear metal cluster required for substrate binding and catalysis. Like other terpenoid cyclases, DCS contains a characteristic aspartate-rich D(307)DTYD(311) motif on helix D that interacts with Mg(2+)(A) and Mg(2+)(C). However, DCS appears to be unique among terpenoid cyclases in that it does not contain the "NSE/DTE" motif on helix H that specifically chelates Mg(2+)(B), which is usually found as the signature sequence (N,D)D(L,I,V)X(S,T)XXXE (boldface indicates Mg(2+)(B) ligands). Instead, DCS contains a second aspartate-rich motif, D(451)DVAE(455), that interacts with Mg(2+)(B). In this regard, DCS is more similar to the isoprenoid chain elongation enzyme farnesyl diphosphate synthase, which also contains two aspartate-rich motifs, rather than the greater family of terpenoid cyclases. Nevertheless, the structure of the DCS-2F-FPP complex shows that the structure of the trinuclear magnesium cluster is generally similar to that of other terpenoid cyclases despite the alternative Mg(2+)(B) binding motif. Analyses of DCS mutants with alanine substitutions in the D(307)DTYD(311) and D(451)DVAE(455) segments reveal the contributions of these segments to catalysis.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, Pennsylvania 19104-6323, USA.
Organizational Affiliation: