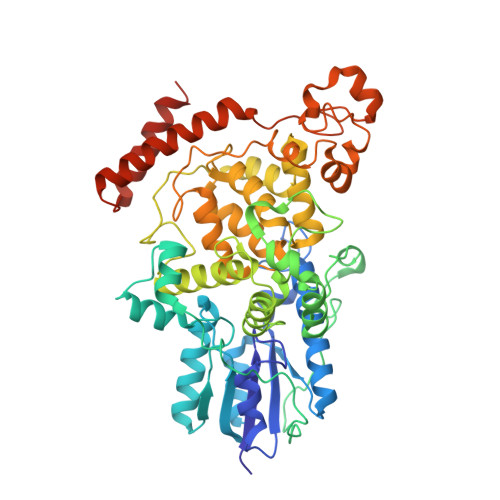
Functional motifs in the (6-4) photolyase crystal structure make a comparative framework for DNA repair photolyases and clock cryptochromes.
Hitomi, K., DiTacchio, L., Arvai, A.S., Yamamoto, J., Kim, S.T., Todo, T., Tainer, J.A., Iwai, S., Panda, S., Getzoff, E.D.(2009) Proc Natl Acad Sci U S A 106: 6962-6967
- PubMed: 19359474 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0809180106
- Primary Citation Related Structures:
3FY4 - PubMed Abstract:
Homologous flavoproteins from the photolyase (PHR)/cryptochrome (CRY) family use the FAD cofactor in PHRs to catalyze DNA repair and in CRYs to tune the circadian clock and control development. To help address how PHR/CRY members achieve these diverse functions, we determined the crystallographic structure of Arabidopsis thaliana (6-4) PHR (UVR3), which is strikingly (>65%) similar in sequence to human circadian clock CRYs. The structure reveals a substrate-binding cavity specific for the UV-induced DNA lesion, (6-4) photoproduct, and cofactor binding sites different from those of bacterial PHRs and consistent with distinct mechanisms for activities and regulation. Mutational analyses were combined with this prototypic structure for the (6-4) PHR/clock CRY cluster to identify structural and functional motifs: phosphate-binding and Pro-Lys-Leu protrusion motifs constricting access to the substrate-binding cavity above FAD, sulfur loop near the external end of the Trp electron-transfer pathway, and previously undefined C-terminal helix. Our results provide a detailed, unified framework for investigations of (6-4) PHRs and the mammalian CRYs. Conservation of key residues and motifs controlling FAD access and activities suggests that regulation of FAD redox properties and radical stability is essential not only for (6-4) photoproduct DNA repair, but also for circadian clock-regulating CRY functions. The structural and functional results reported here elucidate archetypal relationships within this flavoprotein family and suggest how PHRs and CRYs use local residue and cofactor tuning, rather than larger structural modifications, to achieve their diverse functions encompassing DNA repair, plant growth and development, and circadian clock regulation.
- Graduate School of Engineering Science, Osaka University, Toyonaka, Osaka 560-8531, Japan.
Organizational Affiliation: