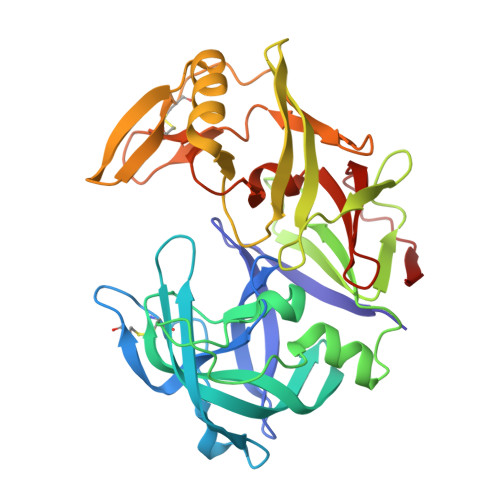
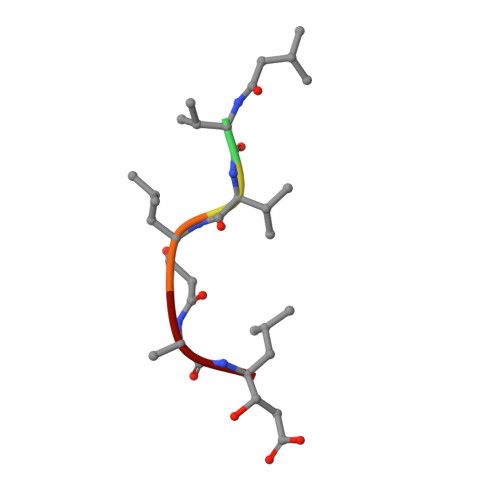
The crystal structure of the secreted aspartic protease 1 from Candida parapsilosis in complex with pepstatin A
Dostal, J., Brynda, J., Hruskova-Heidingsfeldova, O., Sieglova, I., Pichova, I., Rezacova, P.(2009) J Struct Biol 167: 145-152
- PubMed: 19401235 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2009.04.004
- Primary Citation Related Structures:
3FV3 - PubMed Abstract:
Opportunistic pathogens of the genus Candida cause infections representing a major threat to long-term survival of immunocompromised patients. Virulence of the Candida pathogens is enhanced by production of extracellular proteolytic enzymes and secreted aspartic proteases (Saps) are therefore studied as potential virulence factors and possible targets for therapeutic drug design. Candida parapsilosis is less invasive than C. albicans, however, it is one of the leading causative agents of yeast infections. We report three-dimensional crystal structure of Sapp1p from C. parapsilosis in complex with pepstatin A, the classical inhibitor of aspartic proteases. The structure of Sapp1p was determined from protein isolated from its natural source and represents the first structure of Sap from C. parapsilosis. Overall fold and topology of Sapp1p is very similar to the archetypic fold of monomeric aspartic protease family and known structures of Sap isoenzymes from C. albicans and Sapt1p from C. tropicalis. Structural comparison revealed noticeable differences in the structure of loops surrounding the active site. This resulted in differential character, shape, and size of the substrate binding site explaining divergent substrate specificities and inhibitor affinities. Determination of structures of Sap isoenzymes from various species might contribute to the development of new Sap-specific inhibitors.
- Gilead Sciences and IOCB Research Centre, Prague, Czech Republic.
Organizational Affiliation: