Crystal structure of tyrosine aminotransferase tripple mutant (P181Q,R183G,A321K) from Escherichia coli at 2.35 A resolution
Malashkevich, V.N., Ng, B., Kirsch, J.F.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Aromatic-amino-acid aminotransferase | 397 | Escherichia coli K-12 | Mutation(s): 3 Gene Names: b4054, JW4014, tat, tyrB EC: 2.6.1.57 (PDB Primary Data), 2.6.1.107 (UniProt) | 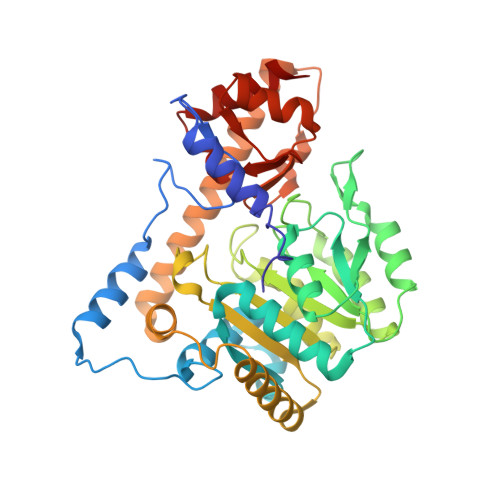 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P04693 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PLR Download:Ideal Coordinates CCD File | G [auth A] H [auth B] I [auth C] J [auth D] K [auth E] | (5-HYDROXY-4,6-DIMETHYLPYRIDIN-3-YL)METHYL DIHYDROGEN PHOSPHATE C8 H12 N O5 P RBCOYOYDYNXAFA-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 98.88 | α = 90 |
| b = 119.16 | β = 90 |
| c = 242.89 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| MOSFLM | data reduction |
| SCALA | data scaling |
| AMoRE | phasing |