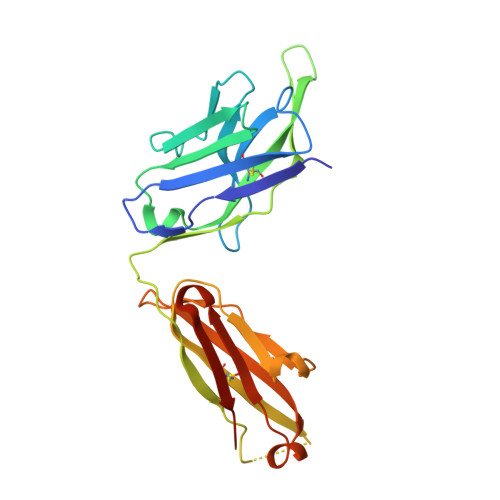
A conformational switch in human immunodeficiency virus gp41 revealed by the structures of overlapping epitopes recognized by neutralizing antibodies.
Pejchal, R., Gach, J.S., Brunel, F.M., Cardoso, R.M., Stanfield, R.L., Dawson, P.E., Burton, D.R., Zwick, M.B., Wilson, I.A.(2009) J Virol 83: 8451-8462
- PubMed: 19515770 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00685-09
- Primary Citation Related Structures:
3FN0 - PubMed Abstract:
The membrane-proximal external region (MPER) of the human immunodeficiency virus (HIV) envelope glycoprotein (gp41) is critical for viral fusion and infectivity and is the target of three of the five known broadly neutralizing HIV type 1 (HIV-1) antibodies, 2F5, Z13, and 4E10. Here, we report the crystal structure of the Fab fragment of Z13e1, an affinity-enhanced variant of monoclonal antibody Z13, in complex with a 12-residue peptide corresponding to the core epitope (W(670)NWFDITN(677)) at 1.8-A resolution. The bound peptide adopts an S-shaped conformation composed of two tandem, perpendicular helical turns. This conformation differs strikingly from the alpha-helical structure adopted by an overlapping MPER peptide bound to 4E10. Z13e1 binds to an elbow in the MPER at the membrane interface, making relatively few interactions with conserved aromatics (Trp672 and Phe673) that are critical for 4E10 recognition. The comparison of the Z13e1 and 4E10 epitope structures reveals a conformational switch such that neutralization can occur by the recognition of the different conformations and faces of the largely amphipathic MPER. The Z13e1 structure provides significant new insights into the dynamic nature of the MPER, which likely is critical for membrane fusion, and it has significant implications for mechanisms of HIV-1 neutralization by MPER antibodies and for the design of HIV-1 immunogens.
- Department of Molecular Biology, The Scripps Research Institute, 10550 N. Torrey Pines Rd., La Jolla, CA 92037, USA.
Organizational Affiliation: