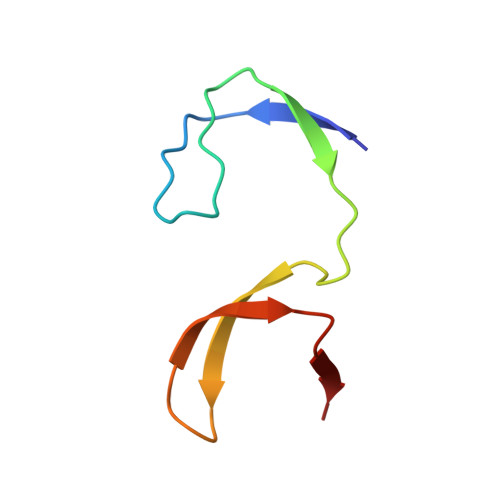
Intertwined dimeric structure for the SH3 domain of the c-Src tyrosine kinase induced by polyethylene glycol binding
Morel, B., Ruiz-Sanz, J., Luque, I.(2009) FEBS Lett 583: 749-753
- PubMed: 19185573 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2009.01.036
- Primary Citation Related Structures:
3FJ5 - PubMed Abstract:
Here we report the first crystal structure of the SH3 domain of the cellular Src tyrosine kinase (c-Src-SH3) domain on its own. In the crystal two molecules of c-Src-SH3 exchange their -RT loops generating an intertwined dimer, in which the two SH3 units, preserving the binding site configuration, are oriented to allow simultaneous binding of two ligand molecules. The dimerization of c-Src-SH3 is induced, both in the crystal and in solution, by the binding of a PEG molecule at the dimer interface, indicating that this type of conformations are energetically close to the native structure. These results have important implications respect to in vivo oligomerization and amyloid aggregation.
- Department of Physical Chemistry, Biochemistry and Inorganic Chemistry, University of Almería, Carretera Sacramento s/n, 04120 Almería, Spain.
Organizational Affiliation: