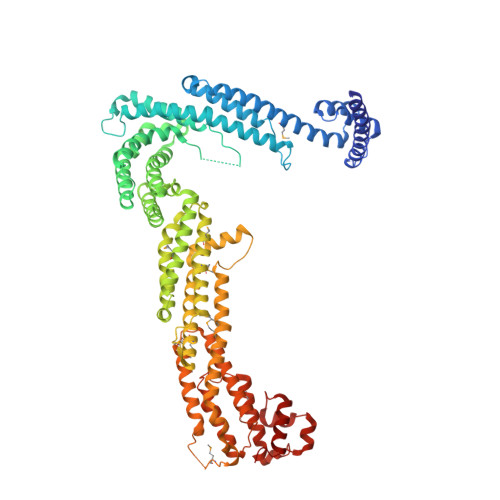
Structural characterization of Tip20p and Dsl1p, subunits of the Dsl1p vesicle tethering complex.
Tripathi, A., Ren, Y., Jeffrey, P.D., Hughson, F.M.(2009) Nat Struct Mol Biol 16: 114-123
- PubMed: 19151722 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1548
- Primary Citation Related Structures:
3ETU, 3ETV, 3FHN - PubMed Abstract:
Multisubunit tethering complexes are essential for intracellular trafficking and have been proposed to mediate the initial interaction between vesicles and the membranes with which they fuse. Here we report initial structural characterization of the Dsl1p complex, whose three subunits are essential for trafficking from the Golgi apparatus to the endoplasmic reticulum (ER). Crystal structures reveal that two of the three subunits, Tip20p and Dsl1p, resemble known subunits of the exocyst complex, establishing a structural connection among several multisubunit tethering complexes and implying that many of their subunits are derived from a common progenitor. We show, moreover, that Tip20p and Dsl1p interact directly via N-terminal alpha-helices. Finally, we establish that different Dsl1p complex subunits bind independently to different ER SNARE proteins. Our results map out two alternative protein-interaction networks capable of tethering COPI-coated vesicles, via the Dsl1p complex, to ER membranes.
- Department of Molecular Biology, Princeton University, Princeton, New Jersey 08544, USA.
Organizational Affiliation: