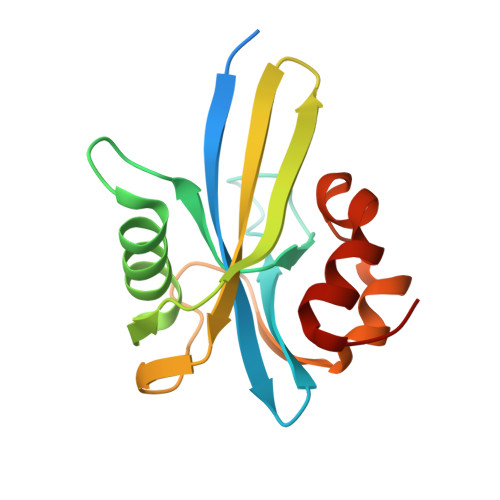
Structure and Biological Function of the RNA Pyrophosphohydrolase BdRppH from Bdellovibrio bacteriovorus.
Messing, S.A., Gabelli, S.B., Liu, Q., Celesnik, H., Belasco, J.G., Pineiro, S.A., Amzel, L.M.(2009) Structure 17: 472-481
- PubMed: 19278661 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2008.12.022
- Primary Citation Related Structures:
3EES, 3EEU, 3EF5, 3FFU - PubMed Abstract:
Until recently, the mechanism of mRNA decay in bacteria was thought to be different from that of eukaryotes. This paradigm changed with the discovery that RppH (ORF176/NudH/YgdP), an Escherichia coli enzyme that belongs to the Nudix superfamily, is an RNA pyrophosphohydrolase that initiates mRNA decay by cleaving pyrophosphate from the 5'-triphosphate. Here we report the 1.9 Angstroms resolution structure of the Nudix hydrolase BdRppH from Bdellovibrio bacteriovorus, a bacterium that feeds on other Gram-negative bacteria. Based on the structure of the enzyme alone and in complex with GTP-Mg2+, we propose a mode of RNA binding similar to that of the nuclear decapping enzyme from Xenopus laevis, X29. In additional experiments, we show that BdRppH can indeed function in vitro and in vivo as an RNA pyrophosphohydrolase. These findings set the basis for the identification of possible decapping enzymes in other bacteria.
- Department of Biophysics and Biophysical Chemistry, School of Medicine, Johns Hopkins University, Baltimore, MD 21205, USA.
Organizational Affiliation: