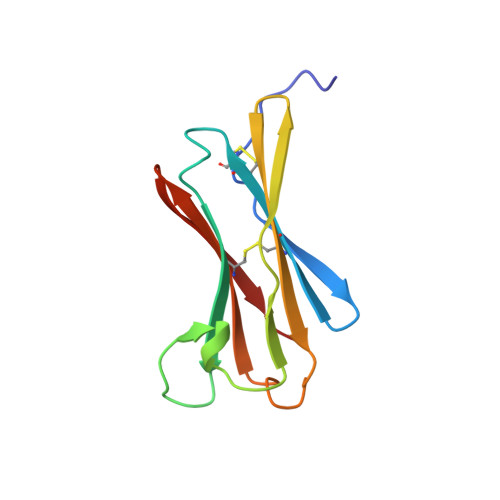
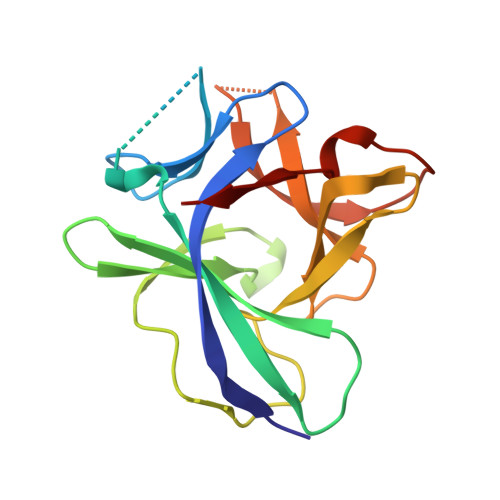
Structural basis for antagonism of human interleukin 18 by poxvirus interleukin 18-binding protein.
Krumm, B., Meng, X., Li, Y., Xiang, Y., Deng, J.(2008) Proc Natl Acad Sci U S A 105: 20711-20715
- PubMed: 19104048 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0809086106
- Primary Citation Related Structures:
3F62 - PubMed Abstract:
Human interleukin-18 (hIL-18) is a cytokine that plays an important role in inflammation and host defense against microbes. Its activity is regulated in vivo by a naturally occurring antagonist, the human IL-18-binding protein (IL-18BP). Functional homologs of human IL-18BP are encoded by all orthopoxviruses, including variola virus, the causative agent of smallpox. They contribute to virulence by suppressing IL-18-mediated immune responses. Here, we describe the 2.0-A resolution crystal structure of an orthopoxvirus IL-18BP, ectromelia virus IL-18BP (ectvIL-18BP), in complex with hIL-18. The hIL-18 structure in the complex shows significant conformational change at the binding interface compared with the structure of ligand-free hIL-18, indicating that the binding is mediated by an induced-fit mechanism. EctvIL-18BP adopts a canonical Ig fold and interacts via one edge of its beta-sandwich with 3 cavities on the hIL-18 surface through extensive hydrophobic and hydrogen bonding interactions. Most of the ectvIL-18BP residues that participate in these interactions are conserved in both human and viral homologs, explaining their functional equivalence despite limited sequence homology. EctvIL-18BP blocks a putative receptor-binding site on IL-18, thus preventing IL-18 from engaging its receptor. Our structure provides insights into how IL-18BPs modulate hIL-18 activity. The revealed binding interface provides the basis for rational design of inhibitors against orthopoxvirus IL-18BP (for treating orthopoxvirus infection) or hIL-18 (for treating certain inflammatory and autoimmune diseases).
- Department of Biochemistry and Molecular Biology, Oklahoma State University, Stillwater, OK 74078, USA.
Organizational Affiliation: