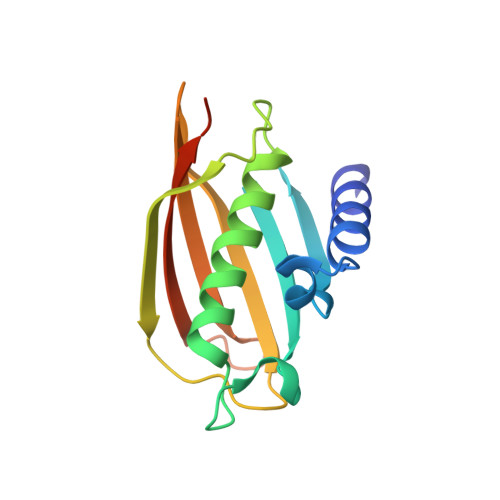
The mechanisms of human hotdog-fold thioesterase 2 (hTHEM2) substrate recognition and catalysis illuminated by a structure and function based analysis
Cao, J., Xu, H., Zhao, H., Gong, W., Dunaway-Mariano, D.(2009) Biochemistry 48: 1293-1304
- PubMed: 19170545 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi801879z
- Primary Citation Related Structures:
3F5O - PubMed Abstract:
The focus of this paper is the hotdog-fold thioesterase THEM2 from human (hTHEM2; Swiss-Prot entry Q9NPJ3 ). In an earlier communication (Cheng, Z., Song, F., Shan, X., Wei, Z., Wang, Y., Dunaway-Mariano, D., and Gong, W. (2006) Crystal structure of human thioesterase superfamily member 2, Biochem. Biophys. Res. Commun. 349, 172-177) we reported the apo crystal structure of hTHEM2. Herein, we report the results of an extensive hTHEM2 substrate screen, the structure determination of hTHEM2 complexed with the inert substrate analogue undecan-2-one-CoA (in which OC-CH(2)-S substitutes for OC-S) and the kinetic analysis of active site mutants. The work described in this paper represents the first reported structure-function based analysis of a human hotdog-fold thioesterase. The catalytic mechanism proposed involves the Asp65/Ser83 assisted attack of a water molecule at the Gly57/Asn50 polarized thioester CO and the Asn50 assisted departure of the thiolate leaving group. Thioesterase activity was observed with acyl-CoAs but not with the human acyl-ACP or with an acyl-Cys peptide. The medium-to-long-chain fatty acyl-CoAs displayed the smallest K(m) values. The substrate specificity profile was analyzed within the context of the liganded enzyme to define the structural determinants of substrate recognition. Based on the results of this structure-function analysis we hypothesize that the physiological role of hTHEM2 involves catalysis of the hydrolysis of cytosolic medium-to-long-chain acyl-CoA thioesters.
- Department of Chemistry and Chemical Biology, University of New Mexico, Albuquerque, New Mexico 87131, USA.
Organizational Affiliation: