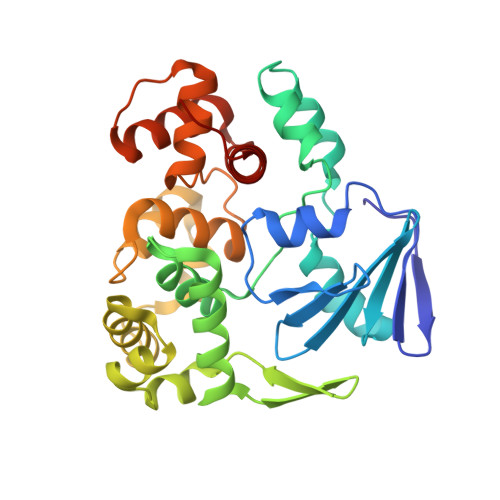
Structural characterization of Clostridium acetobutylicum 8-oxoguanine DNA glycosylase in its apo form and in complex with 8-oxodeoxyguanosine.
Faucher, F., Robey-Bond, S.M., Wallace, S.S., Doublie, S.(2009) J Mol Biology 387: 669-679
- PubMed: 19361427 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.01.067
- Primary Citation Related Structures:
3F0Z, 3F10 - PubMed Abstract:
DNA is subject to a multitude of oxidative damages generated by oxidizing agents from metabolism and exogenous sources and by ionizing radiation. Guanine is particularly vulnerable to oxidation, and the most common oxidative product 8-oxoguanine (8-oxoG) is the most prevalent lesion observed in DNA molecules. 8-OxoG can form a normal Watson-Crick pair with cytosine (8-oxoG:C), but it can also form a stable Hoogsteen pair with adenine (8-oxoG:A), leading to a G:C-->T:A transversion after replication. Fortunately, 8-oxoG is recognized and excised by either of two DNA glycosylases of the base excision repair pathway: formamidopyrimidine-DNA glycosylase and 8-oxoguanine DNA glycosylase (Ogg). While Clostridium acetobutylicum Ogg (CacOgg) DNA glycosylase can specifically recognize and remove 8-oxoG, it displays little preference for the base opposite the lesion, which is unusual for a member of the Ogg1 family. This work describes the crystal structures of CacOgg in its apo form and in complex with 8-oxo-2'-deoxyguanosine. A structural comparison between the apo form and the liganded form of the enzyme reveals a structural reorganization of the C-terminal domain upon binding of 8-oxoG, similar to that reported for human OGG1. A structural comparison of CacOgg with human OGG1, in complex with 8-oxoG containing DNA, provides a structural rationale for the lack of opposite base specificity displayed by CacOgg.
- Department of Microbiology and Molecular Genetics, The Markey Center for Molecular Genetics, University of Vermont, Stafford Hall, 95 Carrigan Drive, Burlington, VT 05405-0068, USA.
Organizational Affiliation: