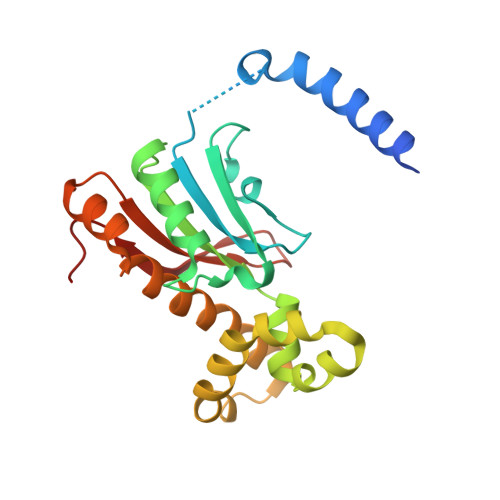
Structural insights into the substrate tunnel of Saccharomyces cerevisiae carbonic anhydrase Nce103.
Teng, Y.B., Jiang, Y.L., He, Y.X., He, W.W., Lian, F.M., Chen, Y., Zhou, C.Z.(2009) BMC Struct Biol 9: 67-67
- PubMed: 19852838 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-9-67
- Primary Citation Related Structures:
3EYX - PubMed Abstract:
The carbonic anhydrases (CAs) are involved in inorganic carbon utilization. They have been classified into six evolutionary and structural families: alpha-, beta-, gamma-, delta-, epsilon-, zeta- CAs, with beta-CAs present in higher plants, algae and prokaryotes. The yeast Saccharomyces cerevisiae encodes a single copy of beta-CA Nce103/YNL036W. We determined the crystal structure of Nce103 in complex with a substrate analog at 2.04 A resolution. It assembles as a homodimer, with the active site located at the interface between two monomers. At the bottom of the substrate pocket, a zinc ion is coordinated by the three highly conserved residues Cys57, His112 and Cys115 in addition to a water molecule. Residues Asp59, Arg61, Gly111, Leu102, Val80, Phe75 and Phe97 form a tunnel to the bottom of the active site which is occupied by a molecule of the substrate analog acetate. Activity assays of full length and two truncated versions of Nce103 indicated that the N-terminal arm is indispensable. The quaternary structure of Nce103 resembles the typical plant type beta-CAs of known structure, with an N-terminal arm indispensable for the enzymatic activity. Comparative structure analysis enables us to draw a possible tunnel for the substrate to access the active site which is located at the bottom of a funnel-shaped substrate pocket.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, 230027, PR China. ybteng@mail.ustc.edu.cn
Organizational Affiliation: