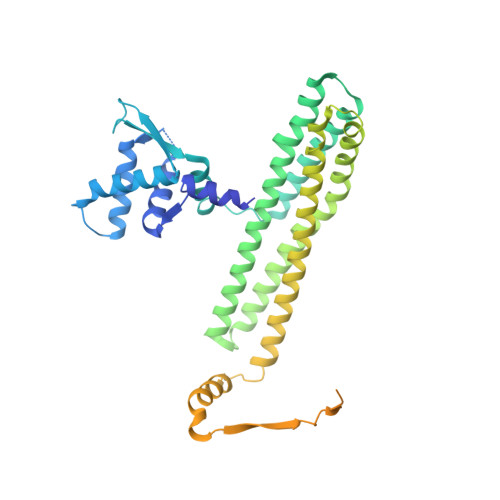
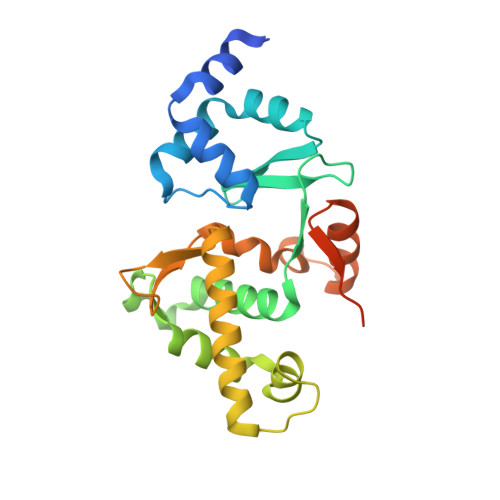
Structural studies of a bacterial condensin complex reveal ATP-dependent disruption of intersubunit interactions.
Woo, J.S., Lim, J.H., Shin, H.C., Suh, M.K., Ku, B., Lee, K.H., Joo, K., Robinson, H., Lee, J., Park, S.Y., Ha, N.C., Oh, B.H.(2009) Cell 136: 85-96
- PubMed: 19135891 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2008.10.050
- Primary Citation Related Structures:
3EUH, 3EUJ, 3EUK - PubMed Abstract:
Condensins are key mediators of chromosome condensation across organisms. Like other condensins, the bacterial MukBEF condensin complex consists of an SMC family protein dimer containing two ATPase head domains, MukB, and two interacting subunits, MukE and MukF. We report complete structural views of the intersubunit interactions of this condensin along with ensuing studies that reveal a role for the ATPase activity of MukB. MukE and MukF together form an elongated dimeric frame, and MukF's C-terminal winged-helix domains (C-WHDs) bind MukB heads to constitute closed ring-like structures. Surprisingly, one of the two bound C-WHDs is forced to detach upon ATP-mediated engagement of MukB heads. This detachment reaction depends on the linker segment preceding the C-WHD, and mutations on the linker restrict cell growth. Thus ATP-dependent transient disruption of the MukB-MukF interaction, which creates openings in condensin ring structures, is likely to be a critical feature of the functional mechanism of condensins.
- Center for Biomolecular Recognition and Division of Molecular and Life Science, Department of Life Sciences, Pohang University of Science and Technology, Pohang, Kyungbuk, 790-784, Korea.
Organizational Affiliation: