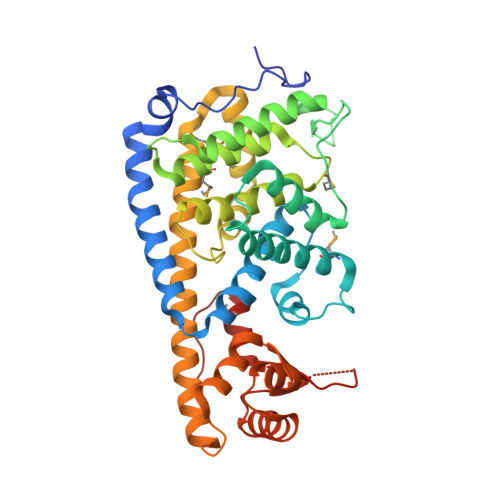
Crystal structure of the Fic (Filamentation induced by cAMP) family protein SO4266 (gi|24375750) from Shewanella oneidensis MR-1 at 1.6 A resolution.
Das, D., Krishna, S.S., McMullan, D., Miller, M.D., Xu, Q., Abdubek, P., Acosta, C., Astakhova, T., Axelrod, H.L., Burra, P., Carlton, D., Chiu, H.J., Clayton, T., Deller, M.C., Duan, L., Elias, Y., Elsliger, M.A., Ernst, D., Feuerhelm, J., Grzechnik, A., Grzechnik, S.K., Hale, J., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Kumar, A., Marciano, D., Morse, A.T., Murphy, K.D., Nigoghossian, E., Okach, L., Oommachen, S., Paulsen, J., Reyes, R., Rife, C.L., Sefcovic, N., Tien, H., Trame, C.B., Trout, C.V., van den Bedem, H., Weekes, D., White, A., Hodgson, K.O., Wooley, J., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2009) Proteins 75: 264-271
- PubMed: 19127588 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.22338
- Primary Citation Related Structures:
3EQX - Joint Center for Structural Genomics, Stanford University, Menlo Park, California, USA.
Organizational Affiliation: