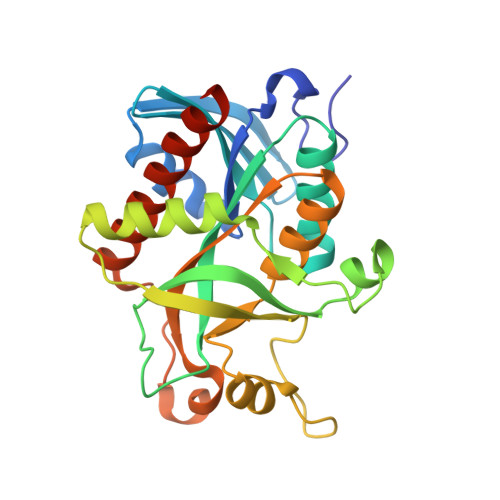
Conservation of structure and activity in Plasmodium purine nucleoside phosphorylases.
Chaikuad, A., Brady, R.L.(2009) BMC Struct Biol 9: 42-42
- PubMed: 19575810 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-9-42
- Primary Citation Related Structures:
3EMV, 3ENZ - PubMed Abstract:
Purine nucleoside phosphorylase (PNP) is central to purine salvage mechanisms in Plasmodium parasites, the causative agents of malaria. Most human malaria results from infection either by Plasmodium falciparum (Pf), the deadliest form of the parasite, or by the widespread Plasmodium vivax (Pv). Whereas the PNP enzyme from Pf has previously been studied in detail, despite the prevalence of Pv little is known about many of the key metabolic enzymes from this parasite, including PvPNP. The crystal structure of PvPNP is described and is seen to have many features in common with the previously reported structure of PfPNP. In particular, the composition and conformations of the active site regions are virtually identical. The crystal structure of a complex of PfPNP co-crystallised with inosine and arsenate is also described, and is found to contain a mixture of products and reactants - hypoxanthine, ribose and arsenate. The ribose C1' in this hybrid complex lies close to the expected point of symmetry along the PNP reaction coordinate, consistent with a conformation between the transition and product states. These two Plasmodium PNP structures confirm the similarity of structure and mechanism of these enzymes, which are also confirmed in enzyme kinetic assays using an array of substrates. These reveal an unusual form of substrate activation by 2'-deoxyinosine of PvPNP, but not PfPNP. The close similarity of the Pf and Pv PNP structures allows characteristic features to be identified that differentiate the Apicomplexa PNPs from the human host enzyme. This similarity also suggests there should be a high level of cross-reactivity for compounds designed to inhibit either of these molecular targets. However, despite these similarities, there are also small differences in the activities of the two Plasmodium enzymes.
- Department of Biochemistry, University of Bristol, Bristol, BS8 1TD, UK. Apirat.Chaikuad@sgc.ox.ac.uk
Organizational Affiliation: