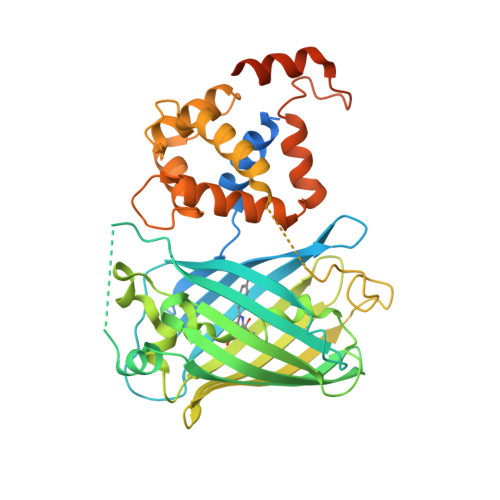
Crystal Structures of the GCaMP Calcium Sensor Reveal the Mechanism of Fluorescence Signal Change and Aid Rational Design
Akerboom, J., Rivera, J.D., Guilbe, M.M., Malave, E.C., Hernandez, H.H., Tian, L., Hires, S.A., Marvin, J.S., Looger, L.L., Schreiter, E.R.(2009) J Biological Chem 284: 6455-6464
- PubMed: 19098007 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M807657200
- Primary Citation Related Structures:
3EK4, 3EK7, 3EK8, 3EKH, 3EKJ - PubMed Abstract:
The genetically encoded calcium indicator GCaMP2 shows promise for neural network activity imaging, but is currently limited by low signal-to-noise ratio. We describe x-ray crystal structures as well as solution biophysical and spectroscopic characterization of GCaMP2 in the calcium-free dark state, and in two calcium-bound bright states: a monomeric form that dominates at intracellular concentrations observed during imaging experiments and an unexpected domain-swapped dimer with decreased fluorescence. This series of structures provides insight into the mechanism of Ca2+-induced fluorescence change. Upon calcium binding, the calmodulin (CaM) domain wraps around the M13 peptide, creating a new domain interface between CaM and the circularly permuted enhanced green fluorescent protein domain. Residues from CaM alter the chemical environment of the circularly permuted enhanced green fluorescent protein chromophore and, together with flexible inter-domain linkers, block solvent access to the chromophore. Guided by the crystal structures, we engineered a series of GCaMP2 point mutants to probe the mechanism of GCaMP2 function and characterized one mutant with significantly improved signal-to-noise. The mutation is located at a domain interface and its effect on sensor function could not have been predicted in the absence of structural data.
- Janelia Farm Research Campus, Howard Hughes Medical Institute, Ashburn, Virginia 20147, USA.
Organizational Affiliation: